A Nanopore Sequencing-based Pharmacogenomic Panel to Personalize Tuberculosis Drug Dosing
- PMID: 38647526
- PMCID: PMC11208962
- DOI: 10.1164/rccm.202309-1583OC
A Nanopore Sequencing-based Pharmacogenomic Panel to Personalize Tuberculosis Drug Dosing
Abstract
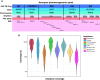
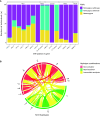
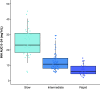
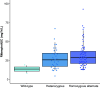
Rationale: Standardized dosing of antitubercular drugs leads to variable plasma drug levels, which are associated with adverse drug reactions, delayed treatment response, and relapse. Mutations in genes affecting drug metabolism explain considerable interindividual pharmacokinetic variability; however, pharmacogenomic assays that predict metabolism of antitubercular drugs have been lacking. Objectives: We sought to develop a Nanopore sequencing panel and validate its performance in patients with active tuberculosis (TB) to personalize treatment dosing. Methods: We developed a Nanopore sequencing panel targeting 15 SNPs in five genes affecting the metabolism of antitubercular drugs. For validation, we sequenced DNA samples (n = 48) from the 1,000 Genomes Project and compared the variant calling accuracy with that of Illumina genome sequencing. We then sequenced DNA samples from patients with active TB (n = 100) from South Africa on a MinION Mk1C and evaluated the relationship between genotypes and pharmacokinetic parameters for isoniazid (INH) and rifampin (RIF). Measurements and Main Results: The pharmacogenomic panel achieved 100% concordance with Illumina sequencing in variant identification for the samples from the 1,000 Genomes Project. In the clinical cohort, coverage was more than 100× for 1,498 of 1,500 (99.8%) amplicons across the 100 samples. Thirty-three percent, 47%, and 20% of participants were identified as slow, intermediate, and rapid INH acetylators, respectively. INH clearance was 2.2 times higher among intermediate acetylators and 3.8 times higher among rapid acetylators, compared with slow acetylators (P < 0.0001). RIF clearance was 17.3% (2.50-29.9) lower in individuals with homozygous AADAC rs1803155 G→A substitutions (P = 0.0015). Conclusions: Targeted sequencing can enable the detection of polymorphisms that influence TB drug metabolism on a low-cost, portable instrument to personalize dosing for TB treatment or prevention.
Keywords: NAT2; Nanopore; pharmacogenomics; targeted sequencing; tuberculosis.
Figures
Update of
-
A Nanopore sequencing-based pharmacogenomic panel to personalize tuberculosis drug dosing.medRxiv [Preprint]. 2023 Sep 10:2023.09.08.23295248. doi: 10.1101/2023.09.08.23295248. medRxiv. 2023. Update in: Am J Respir Crit Care Med. 2024 Jun 15;209(12):1486-1496. doi: 10.1164/rccm.202309-1583OC. PMID: 37732197 Free PMC article. Updated. Preprint.
Comment in
-
Another Step Closer to Genomic-informed Therapeutic Dosing for Tuberculosis Drugs.Am J Respir Crit Care Med. 2024 Jun 15;209(12):1427-1428. doi: 10.1164/rccm.202403-0566ED. Am J Respir Crit Care Med. 2024. PMID: 38648185 Free PMC article. No abstract available.
Similar articles
-
Precision Medicine Strategies to Improve Isoniazid Therapy in Patients with Tuberculosis.Eur J Drug Metab Pharmacokinet. 2024 Sep;49(5):541-557. doi: 10.1007/s13318-024-00910-7. Epub 2024 Aug 17. Eur J Drug Metab Pharmacokinet. 2024. PMID: 39153028 Free PMC article. Review.
-
A Nanopore sequencing-based pharmacogenomic panel to personalize tuberculosis drug dosing.medRxiv [Preprint]. 2023 Sep 10:2023.09.08.23295248. doi: 10.1101/2023.09.08.23295248. medRxiv. 2023. Update in: Am J Respir Crit Care Med. 2024 Jun 15;209(12):1486-1496. doi: 10.1164/rccm.202309-1583OC. PMID: 37732197 Free PMC article. Updated. Preprint.
-
A Rapid Pharmacogenomic Assay to Detect NAT2 Polymorphisms and Guide Isoniazid Dosing for Tuberculosis Treatment.Am J Respir Crit Care Med. 2021 Dec 1;204(11):1317-1326. doi: 10.1164/rccm.202103-0564OC. Am J Respir Crit Care Med. 2021. PMID: 34375564
-
NAT2 genotype guided regimen reduces isoniazid-induced liver injury and early treatment failure in the 6-month four-drug standard treatment of tuberculosis: a randomized controlled trial for pharmacogenetics-based therapy.Eur J Clin Pharmacol. 2013 May;69(5):1091-101. doi: 10.1007/s00228-012-1429-9. Epub 2012 Nov 14. Eur J Clin Pharmacol. 2013. PMID: 23150149 Free PMC article. Clinical Trial.
-
[The role of pharmacogenomics in the tuberculosis treatment regime].Rev Peru Med Exp Salud Publica. 2015 Oct;32(4):794-800. Rev Peru Med Exp Salud Publica. 2015. PMID: 26732931 Review. Spanish.
Cited by
-
Impact of Pharmacogenetics on Pharmacokinetics of First-Line Antituberculosis Drugs in the HIRIF Trial.J Infect Dis. 2025 Aug 14;232(2):e258-e265. doi: 10.1093/infdis/jiaf195. J Infect Dis. 2025. PMID: 40239986 Free PMC article. Clinical Trial.
-
Precision Medicine Strategies to Improve Isoniazid Therapy in Patients with Tuberculosis.Eur J Drug Metab Pharmacokinet. 2024 Sep;49(5):541-557. doi: 10.1007/s13318-024-00910-7. Epub 2024 Aug 17. Eur J Drug Metab Pharmacokinet. 2024. PMID: 39153028 Free PMC article. Review.
-
Genetic and Genomic Approaches to the Study of Drug-Induced Liver Injury.Liver Int. 2025 Jan;45(1):e16191. doi: 10.1111/liv.16191. Liver Int. 2025. PMID: 39704445 Free PMC article. Review.
-
Another Step Closer to Genomic-informed Therapeutic Dosing for Tuberculosis Drugs.Am J Respir Crit Care Med. 2024 Jun 15;209(12):1427-1428. doi: 10.1164/rccm.202403-0566ED. Am J Respir Crit Care Med. 2024. PMID: 38648185 Free PMC article. No abstract available.
-
NAT2 Acetylation Status Predicts Hepatotoxicity During Antituberculosis Therapy: Cumulative Risk Analysis of a Multiethnic Cohort.Int J Mol Sci. 2025 Apr 19;26(8):3881. doi: 10.3390/ijms26083881. Int J Mol Sci. 2025. PMID: 40332508 Free PMC article.
References
-
- Merle CS, Fielding K, Sow OB, Gninafon M, Lo MB, Mthiyane T, et al. OFLOTUB/Gatifloxacin for Tuberculosis Project A four-month gatifloxacin-containing regimen for treating tuberculosis. N Engl J Med . 2014;371:1588–1598. - PubMed
-
- Dorman SE, Nahid P, Kurbatova EV, Goldberg SV, Bozeman L, Burman WJ, et al. AIDS Clinical Trials Group and the Tuberculosis Trials Consortium High-dose rifapentine with or without moxifloxacin for shortening treatment of pulmonary tuberculosis: study protocol for TBTC study 31/ACTG A5349 phase 3 clinical trial. Contemp Clin Trials . 2020;90:105938. - PMC - PubMed
-
- Forget EJ, Menzies D. Adverse reactions to first-line antituberculosis drugs. Expert Opin Drug Saf . 2006;5:231–249. - PubMed
-
- Choi H, Park HA, Hyun IG, Kim JH, Hwang YI, Jang SH, et al. Incidence and outcomes of adverse drug reactions to first-line anti-tuberculosis drugs and their effects on the quality of life: a multicenter prospective cohort study. Pharmacoepidemiol Drug Saf . 2022;31:1153–1163. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

