A viral assembly inhibitor blocks SARS-CoV-2 replication in airway epithelial cells
- PMID: 38649430
- PMCID: PMC11035691
- DOI: 10.1038/s42003-024-06130-8
A viral assembly inhibitor blocks SARS-CoV-2 replication in airway epithelial cells
Abstract
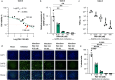
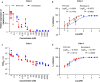
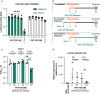
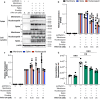
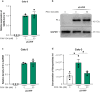
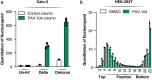
The ongoing evolution of SARS-CoV-2 to evade vaccines and therapeutics underlines the need for innovative therapies with high genetic barriers to resistance. Therefore, there is pronounced interest in identifying new pharmacological targets in the SARS-CoV-2 viral life cycle. The small molecule PAV-104, identified through a cell-free protein synthesis and assembly screen, was recently shown to target host protein assembly machinery in a manner specific to viral assembly. In this study, we investigate the capacity of PAV-104 to inhibit SARS-CoV-2 replication in human airway epithelial cells (AECs). We show that PAV-104 inhibits >99% of infection with diverse SARS-CoV-2 variants in immortalized AECs, and in primary human AECs cultured at the air-liquid interface (ALI) to represent the lung microenvironment in vivo. Our data demonstrate that PAV-104 inhibits SARS-CoV-2 production without affecting viral entry, mRNA transcription, or protein synthesis. PAV-104 interacts with SARS-CoV-2 nucleocapsid (N) and interferes with its oligomerization, blocking particle assembly. Transcriptomic analysis reveals that PAV-104 reverses SARS-CoV-2 induction of the type-I interferon response and the maturation of nucleoprotein signaling pathway known to support coronavirus replication. Our findings suggest that PAV-104 is a promising therapeutic candidate for COVID-19 with a mechanism of action that is distinct from existing clinical management approaches.
© 2024. The Author(s).
Conflict of interest statement
S.S., A.F.L., M.M., S.F.Y. and K.P. are employees of Prosetta Biosciences. V.R.L. is the CEO of Prosetta Biosciences, which manufactures PAV-104 presented in the manuscript. All other authors declare no competing interests.
Figures



Update of
-
A Novel Viral Assembly Inhibitor Blocks SARS-CoV-2 Replication in Airway Epithelial Cells.Res Sq [Preprint]. 2023 May 17:rs.3.rs-2887435. doi: 10.21203/rs.3.rs-2887435/v1. Res Sq. 2023. Update in: Commun Biol. 2024 Apr 22;7(1):486. doi: 10.1038/s42003-024-06130-8. PMID: 37292622 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

