SUCLG1 restricts POLRMT succinylation to enhance mitochondrial biogenesis and leukemia progression
- PMID: 38649537
- PMCID: PMC11183053
- DOI: 10.1038/s44318-024-00101-9
SUCLG1 restricts POLRMT succinylation to enhance mitochondrial biogenesis and leukemia progression
Abstract
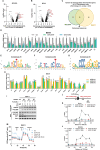
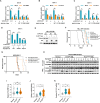
Mitochondria are cellular powerhouses that generate energy through the electron transport chain (ETC). The mitochondrial genome (mtDNA) encodes essential ETC proteins in a compartmentalized manner, however, the mechanism underlying metabolic regulation of mtDNA function remains unknown. Here, we report that expression of tricarboxylic acid cycle enzyme succinate-CoA ligase SUCLG1 strongly correlates with ETC genes across various TCGA cancer transcriptomes. Mechanistically, SUCLG1 restricts succinyl-CoA levels to suppress the succinylation of mitochondrial RNA polymerase (POLRMT). Lysine 622 succinylation disrupts the interaction of POLRMT with mtDNA and mitochondrial transcription factors. SUCLG1-mediated POLRMT hyposuccinylation maintains mtDNA transcription, mitochondrial biogenesis, and leukemia cell proliferation. Specifically, leukemia-promoting FMS-like tyrosine kinase 3 (FLT3) mutations modulate nuclear transcription and upregulate SUCLG1 expression to reduce succinyl-CoA and POLRMT succinylation, resulting in enhanced mitobiogenesis. In line, genetic depletion of POLRMT or SUCLG1 significantly delays disease progression in mouse and humanized leukemia models. Importantly, succinyl-CoA level and POLRMT succinylation are downregulated in FLT3-mutated clinical leukemia samples, linking enhanced mitobiogenesis to cancer progression. Together, SUCLG1 connects succinyl-CoA with POLRMT succinylation to modulate mitochondrial function and cancer development.
Keywords: FMS-like Tyrosine Kinase 3; Lysine Succinylation; Mitochondrial Biogenesis; Mitochondrial RNA Polymerase; Succinate-CoA Ligase.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures










References
-
- Carrozzo R, Verrigni D, Rasmussen M, de Coo R, Amartino H, Bianchi M, Buhas D, Mesli S, Naess K, Born AP, et al. Succinate-CoA ligase deficiency due to mutations in SUCLA2 and SUCLG1: phenotype and genotype correlations in 71 patients. J Inherit Metab Dis. 2016;39:243–252. doi: 10.1007/s10545-015-9894-9. - DOI - PubMed
-
- Chen X, Glytsou C, Zhou H, Narang S, Reyna DE, Lopez A, Sakellaropoulos T, Gong Y, Kloetgen A, Yap YS, et al. Targeting mitochondrial structure sensitizes acute myeloid leukemia to venetoclax treatment. Cancer Discov. 2019;9:890–909. doi: 10.1158/2159-8290.CD-19-0117. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

