Concerted SUMO-targeted ubiquitin ligase activities of TOPORS and RNF4 are essential for stress management and cell proliferation
- PMID: 38649616
- PMCID: PMC11402782
- DOI: 10.1038/s41594-024-01294-7
Concerted SUMO-targeted ubiquitin ligase activities of TOPORS and RNF4 are essential for stress management and cell proliferation
Abstract
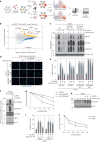
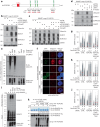
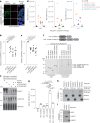
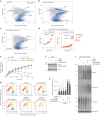
Protein SUMOylation provides a principal driving force for cellular stress responses, including DNA-protein crosslink (DPC) repair and arsenic-induced PML body degradation. In this study, using genome-scale screens, we identified the human E3 ligase TOPORS as a key effector of SUMO-dependent DPC resolution. We demonstrate that TOPORS promotes DPC repair by functioning as a SUMO-targeted ubiquitin ligase (STUbL), combining ubiquitin ligase activity through its RING domain with poly-SUMO binding via SUMO-interacting motifs, analogous to the STUbL RNF4. Mechanistically, TOPORS is a SUMO1-selective STUbL that complements RNF4 in generating complex ubiquitin landscapes on SUMOylated targets, including DPCs and PML, stimulating efficient p97/VCP unfoldase recruitment and proteasomal degradation. Combined loss of TOPORS and RNF4 is synthetic lethal even in unstressed cells, involving defective clearance of SUMOylated proteins from chromatin accompanied by cell cycle arrest and apoptosis. Our findings establish TOPORS as a STUbL whose parallel action with RNF4 defines a general mechanistic principle in crucial cellular processes governed by direct SUMO-ubiquitin crosstalk.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
E1B-55K-Mediated Regulation of RNF4 SUMO-Targeted Ubiquitin Ligase Promotes Human Adenovirus Gene Expression.J Virol. 2018 Jun 13;92(13):e00164-18. doi: 10.1128/JVI.00164-18. Print 2018 Jul 1. J Virol. 2018. PMID: 29695423 Free PMC article.
-
An arginine-rich motif of ring finger protein 4 (RNF4) oversees the recruitment and degradation of the phosphorylated and SUMOylated Krüppel-associated box domain-associated protein 1 (KAP1)/TRIM28 protein during genotoxic stress.J Biol Chem. 2014 Jul 25;289(30):20757-72. doi: 10.1074/jbc.M114.555672. Epub 2014 Jun 6. J Biol Chem. 2014. PMID: 24907272 Free PMC article.
-
RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation.Nat Cell Biol. 2008 May;10(5):538-46. doi: 10.1038/ncb1716. Epub 2008 Apr 13. Nat Cell Biol. 2008. PMID: 18408734
-
SUMO-targeted ubiquitin ligases.Biochim Biophys Acta. 2014 Jan;1843(1):75-85. doi: 10.1016/j.bbamcr.2013.08.022. Epub 2013 Sep 7. Biochim Biophys Acta. 2014. PMID: 24018209 Review.
-
RNF4-A Paradigm for SUMOylation-Mediated Ubiquitination.Proteomics. 2019 Nov;19(21-22):e1900185. doi: 10.1002/pmic.201900185. Epub 2019 Oct 29. Proteomics. 2019. PMID: 31566917 Review.
Cited by
-
Pharmacologic Induction of ERα SUMOylation Disrupts Its Chromatin Binding.ACS Chem Biol. 2024 Nov 15;19(11):2383-2392. doi: 10.1021/acschembio.4c00606. Epub 2024 Oct 21. ACS Chem Biol. 2024. PMID: 39432240 Free PMC article.
-
Multifaceted roles of SUMO in DNA metabolism.Nucleus. 2024 Dec;15(1):2398450. doi: 10.1080/19491034.2024.2398450. Epub 2024 Sep 17. Nucleus. 2024. PMID: 39287196 Free PMC article. Review.
-
PML mutants from arsenic-resistant patients reveal SUMO1-TOPORS and SUMO2/3-RNF4 degradation pathways.J Cell Biol. 2025 Jun 2;224(6):e202407133. doi: 10.1083/jcb.202407133. Epub 2025 Apr 16. J Cell Biol. 2025. PMID: 40239066 Free PMC article.
-
Allosteric activation of the SPRTN protease by ubiquitin maintains genome stability.Nat Commun. 2025 Jul 21;16(1):5422. doi: 10.1038/s41467-025-61224-z. Nat Commun. 2025. PMID: 40691134 Free PMC article.
-
Immune regulation by the SUMO family.Nat Rev Immunol. 2025 Aug;25(8):608-620. doi: 10.1038/s41577-025-01155-4. Epub 2025 Mar 19. Nat Rev Immunol. 2025. PMID: 40108400 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

