Cortical gene expression architecture links healthy neurodevelopment to the imaging, transcriptomics and genetics of autism and schizophrenia
- PMID: 38649755
- PMCID: PMC11156586
- DOI: 10.1038/s41593-024-01624-4
Cortical gene expression architecture links healthy neurodevelopment to the imaging, transcriptomics and genetics of autism and schizophrenia
Abstract
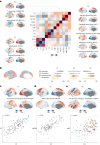
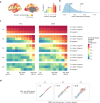
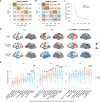
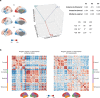
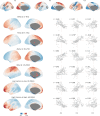
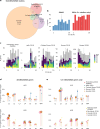
Human brain organization involves the coordinated expression of thousands of genes. For example, the first principal component (C1) of cortical transcription identifies a hierarchy from sensorimotor to association regions. In this study, optimized processing of the Allen Human Brain Atlas revealed two new components of cortical gene expression architecture, C2 and C3, which are distinctively enriched for neuronal, metabolic and immune processes, specific cell types and cytoarchitectonics, and genetic variants associated with intelligence. Using additional datasets (PsychENCODE, Allen Cell Atlas and BrainSpan), we found that C1-C3 represent generalizable transcriptional programs that are coordinated within cells and differentially phased during fetal and postnatal development. Autism spectrum disorder and schizophrenia were specifically associated with C1/C2 and C3, respectively, across neuroimaging, differential expression and genome-wide association studies. Evidence converged especially in support of C3 as a normative transcriptional program for adolescent brain development, which can lead to atypical supragranular cortical connectivity in people at high genetic risk for schizophrenia.
© 2024. The Author(s).
Conflict of interest statement
K.M.A. is an employee of Neumora Therapeutics. R.D.M. is an employee of Octave Biosciences. E.T.B. has consulted for Boehringer Ingelheim, SR One, GlaxoSmithKline, Sosei Heptares and Monument Therapeutics. All other authors have no disclosures to make.
Figures




References
-
- van den Heuvel MP, Yeo BTT. A spotlight on bridging microscale and macroscale human brain architecture. Neuron. 2017;93:1248–1251. - PubMed
MeSH terms
Grants and funding
- T32 MH019112/MH/NIMH NIH HHS/United States
- MR/M009041/1/MRC_/Medical Research Council/United Kingdom
- K08 MH120564/MH/NIMH NIH HHS/United States
- K08MH120564/U.S. Department of Health & Human Services | NIH | National Institute of Mental Health (NIMH)
- 1ZIAMH002949-04/U.S. Department of Health & Human Services | NIH | National Institute of Mental Health (NIMH)
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

