Proteomic studies in VWA1-related neuromyopathy allowed new pathophysiological insights and the definition of blood biomarkers
- PMID: 38652110
- PMCID: PMC11037410
- DOI: 10.1111/jcmm.18122
Proteomic studies in VWA1-related neuromyopathy allowed new pathophysiological insights and the definition of blood biomarkers
Abstract
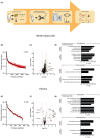
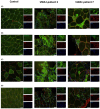
Bi-allelic variants in VWA1, encoding Von Willebrand Factor A domain containing 1 protein localized to the extracellular matrix (ECM), were linked to a neuromuscular disorder with manifestation in child- or adulthood. Clinical findings indicate a neuromyopathy presenting with muscle weakness. Given that pathophysiological processes are still incompletely understood, and biomarkers are still missing, we aimed to identify blood biomarkers of pathophysiological relevance: white blood cells (WBC) and plasma derived from six VWA1-patients were investigated by proteomics. Four proteins, BET1, HNRNPDL, NEFM and PHGDH, known to be involved in neurological diseases and dysregulated in WBC were further validated by muscle-immunostainings unravelling HNRNPDL as a protein showing differences between VWA1-patients, healthy controls and patients suffering from neurogenic muscular atrophy and BICD2-related neuromyopathy. Immunostaining studies of PHGDH indicate its involvement in apoptotic processes via co-localisation with caspase-3. NEFM showed an increase in cells within the ECM in biopsies of all patients studied. Plasma proteomics unravelled dysregulation of 15 proteins serving as biomarker candidates among which a profound proportion of increased ones (6/11) are mostly related to antioxidative processes and have even partially been described as blood biomarkers for other entities of neuromuscular disorders before. CRP elevated in plasma also showed an increase in the extracellular space of VWA1-mutant muscle. Results of our combined studies for the first time describe pathophysiologically relevant biomarkers for VWA1-related neuromyopathy and suggest that VWA1-patient derived blood might hold the potential to study disease processes of clinical relevance, an important aspect for further preclinical studies.
Keywords: BET1; HNRNPDL; NEFM and PHGDH; Von Willebrand factor a domain containing 1 protein; neuromyopathy.
© 2024 The Authors. Journal of Cellular and Molecular Medicine published by Foundation for Cellular and Molecular Medicine and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare to not have any conflict of interest.
Figures

Similar articles
-
Autosomal recessive VWA1-related disorder: comprehensive analysis of phenotypic variability and genetic mutations.Brain Commun. 2024 Oct 28;6(6):fcae377. doi: 10.1093/braincomms/fcae377. eCollection 2024. Brain Commun. 2024. PMID: 39502942 Free PMC article.
-
Bi-allelic truncating mutations in VWA1 cause neuromyopathy.Brain. 2021 Mar 3;144(2):574-583. doi: 10.1093/brain/awaa418. Brain. 2021. PMID: 33459760
-
Upper motor neuron signs and early onset gait abnormalities in young children with bi-allelic VWA1 variants.Am J Med Genet A. 2022 Dec;188(12):3531-3534. doi: 10.1002/ajmg.a.62953. Epub 2022 Aug 17. Am J Med Genet A. 2022. PMID: 35975723
-
Proteomic serum biomarkers for neuromuscular diseases.Expert Rev Proteomics. 2018 Mar;15(3):277-291. doi: 10.1080/14789450.2018.1429923. Epub 2018 Jan 23. Expert Rev Proteomics. 2018. PMID: 29338453 Review.
-
Proteomic Profiling Towards a Better Understanding of Genetic Based Muscular Diseases: The Current Picture and a Look to the Future.Biomolecules. 2025 Jan 15;15(1):130. doi: 10.3390/biom15010130. Biomolecules. 2025. PMID: 39858524 Free PMC article. Review.
Cited by
-
Autosomal recessive VWA1-related disorder: comprehensive analysis of phenotypic variability and genetic mutations.Brain Commun. 2024 Oct 28;6(6):fcae377. doi: 10.1093/braincomms/fcae377. eCollection 2024. Brain Commun. 2024. PMID: 39502942 Free PMC article.
References
-
- Bhatt JM. The epidemiology of neuromuscular diseases. Neurol Clin. 2016;34(4):999‐1021. - PubMed
-
- Thompson R, Spendiff S, Roos A, et al. Advances in the diagnosis of inherited neuromuscular diseases and implications for therapy development. Lancet Neurol. 2020;19(6):522‐532. - PubMed
-
- Arribat Y. Genetic alterations of VWA1: a new link between extracellular matrix and neuromuscular diseases. Brain. 2021;144(2):362‐365. - PubMed
-
- Legay C, Dobbertin A. Collagens at the vertebrate neuromuscular junction, from structure to pathologies. Neurosci Lett. 2020;735:135155. - PubMed
-
- Deschauer M, Hengel H, Rupprich K, et al. Bi‐allelic truncating mutations in VWA1 cause neuromyopathy. Brain. 2021;144(2):574‐583. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

