Deciphering Staphylococcus aureus- host dynamics using dual activity-based protein profiling of ATP-interacting proteins
- PMID: 38656122
- PMCID: PMC11097646
- DOI: 10.1128/msystems.00179-24
Deciphering Staphylococcus aureus- host dynamics using dual activity-based protein profiling of ATP-interacting proteins
Abstract
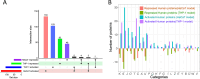
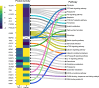
The utilization of ATP within cells plays a fundamental role in cellular processes that are essential for the regulation of host-pathogen dynamics and the subsequent immune response. This study focuses on ATP-binding proteins to dissect the complex interplay between Staphylococcus aureus and human cells, particularly macrophages (THP-1) and keratinocytes (HaCaT), during an intracellular infection. A snapshot of the various protein activity and function is provided using a desthiobiotin-ATP probe, which targets ATP-interacting proteins. In S. aureus, we observe enrichment in pathways required for nutrient acquisition, biosynthesis and metabolism of amino acids, and energy metabolism when located inside human cells. Additionally, the direct profiling of the protein activity revealed specific adaptations of S. aureus to the keratinocytes and macrophages. Mapping the differentially activated proteins to biochemical pathways in the human cells with intracellular bacteria revealed cell-type-specific adaptations to bacterial challenges where THP-1 cells prioritized immune defenses, autophagic cell death, and inflammation. In contrast, HaCaT cells emphasized barrier integrity and immune activation. We also observe bacterial modulation of host processes and metabolic shifts. These findings offer valuable insights into the dynamics of S. aureus-host cell interactions, shedding light on modulating host immune responses to S. aureus, which could involve developing immunomodulatory therapies.
Importance: This study uses a chemoproteomic approach to target active ATP-interacting proteins and examines the dynamic proteomic interactions between Staphylococcus aureus and human cell lines THP-1 and HaCaT. It uncovers the distinct responses of macrophages and keratinocytes during bacterial infection. S. aureus demonstrated a tailored response to the intracellular environment of each cell type and adaptation during exposure to professional and non-professional phagocytes. It also highlights strategies employed by S. aureus to persist within host cells. This study offers significant insights into the human cell response to S. aureus infection, illuminating the complex proteomic shifts that underlie the defense mechanisms of macrophages and keratinocytes. Notably, the study underscores the nuanced interplay between the host's metabolic reprogramming and immune strategy, suggesting potential therapeutic targets for enhancing host defense and inhibiting bacterial survival. The findings enhance our understanding of host-pathogen interactions and can inform the development of targeted therapies against S. aureus infections.
Keywords: ATP-interacting proteins; HaCaT cells; Staphylococcus aureus; THP-1 cells; activity-based protein profiling (ABPP); bacterial metabolism; host immune response; host–pathogen interactions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures







Similar articles
-
Mapping of interactions between human macrophages and Staphylococcus aureus reveals an involvement of MAP kinase signaling in the host defense.J Proteome Res. 2011 Sep 2;10(9):4018-32. doi: 10.1021/pr200224x. Epub 2011 Jul 28. J Proteome Res. 2011. PMID: 21736355
-
Type VII secretion system extracellular protein B targets STING to evade host anti-Staphylococcus aureus immunity.Proc Natl Acad Sci U S A. 2024 May 28;121(22):e2402764121. doi: 10.1073/pnas.2402764121. Epub 2024 May 21. Proc Natl Acad Sci U S A. 2024. PMID: 38771879 Free PMC article.
-
Consequences of Metabolic Interactions during Staphylococcus aureus Infection.Toxins (Basel). 2020 Sep 9;12(9):581. doi: 10.3390/toxins12090581. Toxins (Basel). 2020. PMID: 32917040 Free PMC article. Review.
-
Endogenous nitric oxide promotes Staphylococcus aureus virulence by activating autophagy.mBio. 2025 Apr 9;16(4):e0400624. doi: 10.1128/mbio.04006-24. Epub 2025 Feb 25. mBio. 2025. PMID: 39998210 Free PMC article.
-
Epic Immune Battles of History: Neutrophils vs. Staphylococcus aureus.Front Cell Infect Microbiol. 2017 Jun 30;7:286. doi: 10.3389/fcimb.2017.00286. eCollection 2017. Front Cell Infect Microbiol. 2017. PMID: 28713774 Free PMC article. Review.
References
MeSH terms
LinkOut - more resources
Full Text Sources

