Detection and identification of single ribonucleotide monophosphates using a dual in-plane nanopore sensor made in a thermoplastic via replication
- PMID: 38656267
- PMCID: PMC11091956
- DOI: 10.1039/d3lc01062g
Detection and identification of single ribonucleotide monophosphates using a dual in-plane nanopore sensor made in a thermoplastic via replication
Abstract
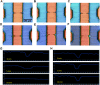
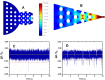
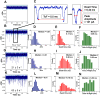
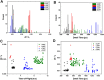
We report the generation of ∼8 nm dual in-plane pores fabricated in a thermoplastic via nanoimprint lithography (NIL). These pores were connected in series with nanochannels, one of which served as a flight tube to allow the identification of single molecules based on their molecular-dependent apparent mobilities (i.e., dual in-plane nanopore sensor). Two different thermoplastics were investigated including poly(methyl methacrylate), PMMA, and cyclic olefin polymer, COP, as the substrate for the sensor both of which were sealed using a low glass transition cover plate (cyclic olefin co-polymer, COC) that could be thermally fusion bonded to the PMMA or COP substrate at a temperature minimizing nanostructure deformation. Unique to these dual in-plane nanopore sensors was two pores flanking each side of the nanometer flight tube (50 × 50 nm, width × depth) that was 10 μm in length. The utility of this dual in-plane nanopore sensor was evaluated to not only detect, but also identify single ribonucleotide monophosphates (rNMPs) by using the travel time (time-of-flight, ToF), the resistive pulse event amplitude, and the dwell time. In spite of the relatively large size of these in-plane pores (∼8 nm effective diameter), we could detect via resistive pulse sensing (RPS) single rNMP molecules at a mass load of 3.9 fg, which was ascribed to the unique structural features of the nanofluidic network and the use of a thermoplastic with low relative dielectric constants, which resulted in a low RMS noise level in the open pore current. Our data indicated that the identification accuracy of individual rNMPs was high, which was ascribed to an improved chromatographic contribution to the nano-electrophoresis apparent mobility. With the ToF data only, the identification accuracy was 98.3%. However, when incorporating the resistive pulse sensing event amplitude and dwell time in conjunction with the ToF and analyzed via principal component analysis (PCA), the identification accuracy reached 100%. These findings pave the way for the realization of a novel chip-based single-molecule RNA sequencing technology.
Conflict of interest statement
The authors have declared no conflict of interest.
Figures



References
-
- Corney D. C. Mater. Methods. 2013;3:203.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

