AmplificationTimeR: an R package for timing sequential amplification events
- PMID: 38656989
- PMCID: PMC11153944
- DOI: 10.1093/bioinformatics/btae281
AmplificationTimeR: an R package for timing sequential amplification events
Abstract
Motivation: Few methods exist for timing individual amplification events in regions of focal amplification. Current methods are also limited in the copy number states that they are able to time. Here we introduce AmplificationTimeR, a method for timing higher level copy number gains and inferring the most parsimonious order of events for regions that have undergone both single gains and whole genome duplication. Our method is an extension of established approaches for timing genomic gains.
Results: We can time more copy number states, and in states covered by other methods our results are comparable to previously published methods.
Availability and implementation: AmplificationTimer is freely available as an R package hosted at https://github.com/Wedge-lab/AmplificationTimeR.
© The Author(s) 2024. Published by Oxford University Press.
Conflict of interest statement
None declared.
Figures


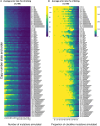

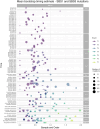
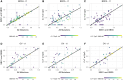
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

