Microbiota metabolism of intestinal amino acids impacts host nutrient homeostasis and physiology
- PMID: 38657606
- PMCID: PMC11636940
- DOI: 10.1016/j.chom.2024.04.004
Microbiota metabolism of intestinal amino acids impacts host nutrient homeostasis and physiology
Abstract
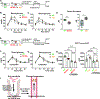
The intestine and liver are thought to metabolize dietary nutrients and regulate host nutrient homeostasis. Here, we find that the gut microbiota also reshapes the host amino acid (aa) landscape via efficiently metabolizing intestinal aa. To identify the responsible microbes/genes, we developed a metabolomics-based assay to screen 104 commensals and identified candidates that efficiently utilize aa. Using genetics, we identified multiple responsible metabolic genes in phylogenetically diverse microbes. By colonizing germ-free mice with the wild-type strain and their isogenic mutant deficient in individual aa-metabolizing genes, we found that these genes regulate the availability of gut and circulatory aa. Notably, microbiota genes for branched-chain amino acids (BCAAs) and tryptophan metabolism indirectly affect host glucose homeostasis via peripheral serotonin. Collectively, at single-gene level, this work characterizes a microbiota-encoded metabolic activity that affects host nutrient homeostasis and provides a roadmap to interrogate microbiota-dependent activity to improve human health.
Keywords: amino acid metabolism; glucose tolerance; gut microbiota and metabolic genes; human microbiota.
Copyright © 2024 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests D.A. has contributed to scientific advisory boards at Pfizer, Takeda, and the Kenneth Rainin Foundation.
Figures


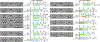

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

