Discovery of WRN inhibitor HRO761 with synthetic lethality in MSI cancers
- PMID: 38658754
- PMCID: PMC11078746
- DOI: 10.1038/s41586-024-07350-y
Discovery of WRN inhibitor HRO761 with synthetic lethality in MSI cancers
Abstract
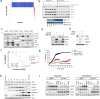
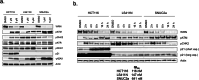
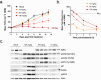
The Werner syndrome RecQ helicase WRN was identified as a synthetic lethal target in cancer cells with microsatellite instability (MSI) by several genetic screens1-6. Despite advances in treatment with immune checkpoint inhibitors7-10, there is an unmet need in the treatment of MSI cancers11-14. Here we report the structural, biochemical, cellular and pharmacological characterization of the clinical-stage WRN helicase inhibitor HRO761, which was identified through an innovative hit-finding and lead-optimization strategy. HRO761 is a potent, selective, allosteric WRN inhibitor that binds at the interface of the D1 and D2 helicase domains, locking WRN in an inactive conformation. Pharmacological inhibition by HRO761 recapitulated the phenotype observed by WRN genetic suppression, leading to DNA damage and inhibition of tumour cell growth selectively in MSI cells in a p53-independent manner. Moreover, HRO761 led to WRN degradation in MSI cells but not in microsatellite-stable cells. Oral treatment with HRO761 resulted in dose-dependent in vivo DNA damage induction and tumour growth inhibition in MSI cell- and patient-derived xenograft models. These findings represent preclinical pharmacological validation of WRN as a therapeutic target in MSI cancers. A clinical trial with HRO761 (NCT05838768) is ongoing to assess the safety, tolerability and preliminary anti-tumour activity in patients with MSI colorectal cancer and other MSI solid tumours.
© 2024. The Author(s).
Conflict of interest statement
M.C.-C., H.V., C.Q., L.L., J.S., M.W., E.B., F.Z., E.G., E.D., S.W., S.B. and F.S. are employees of Novartis Pharma and hold stock in Novartis, Alcon and Sandoz. H.M., R.d.K., J.B., M.R., N.D.N., C.S., J. Hamon, M.S., A.H., S.F., M.H., R.A.-R., G.K., A.D., M.D., C.H., V.R., H.-J.M. and V.B. are employees of Novartis Pharma and hold stock in Novartis and Sandoz. I.J. and J. Hinrichs are employees of Novartis Pharma and hold stock in Novartis. R.S. is an employee of Novartis Pharma. F.H. and E.J.N. are employees of Pierre Fabre and hold stock in Novartis. HRO761 is covered by patent WO2022/249060 with the following co-authors listed as inventors: V.B., A.D., J. Hamon, J. Hinrichs, H.M., J.S., R.S. and F.Z. Additional patent applications related to WO2022/249060 are pending.
Figures









References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

