Single-cell analysis reveals context-dependent, cell-level selection of mtDNA
- PMID: 38658765
- PMCID: PMC11078733
- DOI: 10.1038/s41586-024-07332-0
Single-cell analysis reveals context-dependent, cell-level selection of mtDNA
Abstract
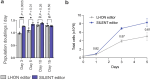
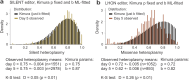
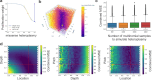
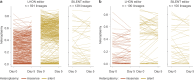
Heteroplasmy occurs when wild-type and mutant mitochondrial DNA (mtDNA) molecules co-exist in single cells1. Heteroplasmy levels change dynamically in development, disease and ageing2,3, but it is unclear whether these shifts are caused by selection or drift, and whether they occur at the level of cells or intracellularly. Here we investigate heteroplasmy dynamics in dividing cells by combining precise mtDNA base editing (DdCBE)4 with a new method, SCI-LITE (single-cell combinatorial indexing leveraged to interrogate targeted expression), which tracks single-cell heteroplasmy with ultra-high throughput. We engineered cells to have synonymous or nonsynonymous complex I mtDNA mutations and found that cell populations in standard culture conditions purge nonsynonymous mtDNA variants, whereas synonymous variants are maintained. This suggests that selection dominates over simple drift in shaping population heteroplasmy. We simultaneously tracked single-cell mtDNA heteroplasmy and ancestry, and found that, although the population heteroplasmy shifts, the heteroplasmy of individual cell lineages remains stable, arguing that selection acts at the level of cell fitness in dividing cells. Using these insights, we show that we can force cells to accumulate high levels of truncating complex I mtDNA heteroplasmy by placing them in environments where loss of biochemical complex I activity has been reported to benefit cell fitness. We conclude that in dividing cells, a given nonsynonymous mtDNA heteroplasmy can be harmful, neutral or even beneficial to cell fitness, but that the 'sign' of the effect is wholly dependent on the environment.
© 2024. The Author(s).
Conflict of interest statement
V.K.M. is a consultant to 5am Ventures. All other authors declare no competing interests.
Figures










References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

