This is a preprint.
Prophage-encoded methyltransferase drives adaptation of community-acquired methicillin-resistant Staphylococcus aureus
- PMID: 38659881
- PMCID: PMC11042277
- DOI: 10.1101/2024.04.17.589803
Prophage-encoded methyltransferase drives adaptation of community-acquired methicillin-resistant Staphylococcus aureus
Update in
-
Prophage-encoded methyltransferase drives adaptation of community-acquired methicillin-resistant Staphylococcus aureus.J Clin Invest. 2025 Jul 22;135(18):e177872. doi: 10.1172/JCI177872. eCollection 2025 Sep 16. J Clin Invest. 2025. PMID: 40700354 Free PMC article.
Abstract
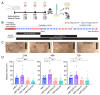
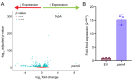
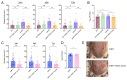
We recently described the evolution of a community-acquired methicillin-resistant Staphylococcus aureus (CA-MRSA) USA300 variant responsible for an outbreak of skin and soft tissue infections. Acquisition of a mosaic version of the Φ11 prophage (mΦ11) that increases skin abscess size was an early step in CA-MRSA adaptation that primed the successful spread of the clone. The present report shows how prophage mΦ11 exerts its effect on virulence for skin infection without encoding a known toxin or fitness genes. Abscess size and skin inflammation were associated with DNA methylase activity of an mΦ11-encoded adenine methyltransferase (designated pamA). pamA increased expression of fibronectin-binding protein A (fnbA; FnBPA), and inactivation of fnbA eliminated the effect of pamA on abscess virulence without affecting strains lacking pamA. Thus, fnbA is a pamA-specific virulence factor. Mechanistically, pamA was shown to promote biofilm formation in vivo in skin abscesses, a phenotype linked to FnBPA's role in biofilm formation. Collectively, these data reveal a novel mechanism-epigenetic regulation of staphylococcal gene expression-by which phage can regulate virulence to drive adaptive leaps by S. aureus.
Conflict of interest statement
Conflict of Interest Statement: V.J.T. has received honoraria from Pfizer and MedImmune, and is an inventor on patents and patent applications filed by New York University, which are currently under commercial license to Janssen Biotech Inc. He is currently an advisor for Moderna. All other authors: no competing interests declared.
Figures




References
-
- Weber JT. Community-associated methicillin-resistant Staphylococcus aureus. Clin Infect Dis. 2005;41 Suppl 4:S269–72. - PubMed
-
- Herold BC, Immergluck LC, Maranan MC, Lauderdale DS, Gaskin RE, Boyle-Vavra S, et al. Community-acquired methicillin-resistant Staphylococcus aureus in children with no identified predisposing risk. JAMA. 1998;279(8):593–8. - PubMed
-
- Moran GJ, Krishnadasan A, Gorwitz RJ, Fosheim GE, McDougal LK, Carey RB, et al. Methicillin-resistant S. aureus infections among patients in the emergency department. N Engl J Med. 2006;355(7):666–74. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
