Chloroplast genome structure analysis of Equisetum unveils phylogenetic relationships to ferns and mutational hotspot region
- PMID: 38665369
- PMCID: PMC11044155
- DOI: 10.3389/fpls.2024.1328080
Chloroplast genome structure analysis of Equisetum unveils phylogenetic relationships to ferns and mutational hotspot region
Abstract
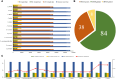
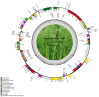
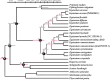
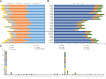
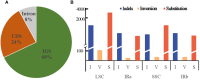
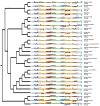
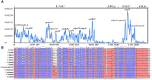
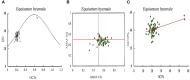
Equisetum is one of the oldest extant group vascular plants and is considered to be the key to understanding vascular plant evolution. Equisetum is distributed almost all over the world and has a high degree of adaptability to different environments. Despite the fossil record of horsetails (Equisetum, Equisetaceae) dating back to the Carboniferous, the phylogenetic relationship of this genus is not well, and the chloroplast evolution in Equisetum remains poorly understood. In order to fill this gap, we sequenced, assembled, and annotated the chloroplast genomes of 12 species of Equisetum, and compared them to 13 previously published vascular plants chloroplast genomes to deeply examine the plastome evolutionary dynamics of Equisetum. The chloroplast genomes have a highly conserved quadripartite structure across the genus, but these chloroplast genomes have a lower GC content than other ferns. The size of Equisetum plastomes ranges from 130,773 bp to 133,684 bp and they encode 130 genes. Contraction/expansion of IR regions and the number of simple sequences repeat regions underlie large genomic variations in size among them. Comparative analysis revealed we also identified 13 divergence hotspot regions. Additionally, the genes accD and ycf1 can be used as potential DNA barcodes for the identification and phylogeny of the genus Equisetum. Twelve photosynthesis-related genes were specifically selected in Equisetum. Comparative genomic analyses implied divergent evolutionary patterns between Equisetum and other ferns. Phylogenomic analyses and molecular dating revealed a relatively distant phylogenetic relationship between Equisetum and other ferns, supporting the division of pteridophyte into Lycophytes, Equisetaceae and ferns. The results show that the chloroplast genome can be used to solve phylogenetic problems within or between Equisetum species, and also provide genomic resources for the study of Equisetum systematics and evolution.
Keywords: divergent hotspot; evolutionary; phylogenomics; pteridophytes; sequence characteristic.
Copyright © 2024 Sun, Wei, Gu, Wang, Liu and Yan.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Biogeography and genome size evolution of the oldest extant vascular plant genus, Equisetum (Equisetaceae).Ann Bot. 2021 Apr 17;127(5):681-695. doi: 10.1093/aob/mcab005. Ann Bot. 2021. PMID: 33598697 Free PMC article.
-
Plastome phylogenomics unravels the evolutionary relationships and biogeographic history of Chloranthaceae.BMC Plant Biol. 2025 Apr 25;25(1):543. doi: 10.1186/s12870-025-06586-8. BMC Plant Biol. 2025. PMID: 40281417 Free PMC article.
-
Complete chloroplast genome sequence of a tree fern Alsophila spinulosa: insights into evolutionary changes in fern chloroplast genomes.BMC Evol Biol. 2009 Jun 11;9:130. doi: 10.1186/1471-2148-9-130. BMC Evol Biol. 2009. PMID: 19519899 Free PMC article.
-
Complete plastid genomes from Ophioglossum californicum, Psilotum nudum, and Equisetum hyemale reveal an ancestral land plant genome structure and resolve the position of Equisetales among monilophytes.BMC Evol Biol. 2013 Jan 11;13:8. doi: 10.1186/1471-2148-13-8. BMC Evol Biol. 2013. PMID: 23311954 Free PMC article.
-
Trends and concepts in fern classification.Ann Bot. 2014 Mar;113(4):571-94. doi: 10.1093/aob/mct299. Epub 2014 Feb 13. Ann Bot. 2014. PMID: 24532607 Free PMC article. Review.
Cited by
-
The first complete mitochondrial genome assembly and comparative analysis of the fern Blechnaceae family: Blechnopsis orientalis.Front Plant Sci. 2025 Mar 20;16:1534171. doi: 10.3389/fpls.2025.1534171. eCollection 2025. Front Plant Sci. 2025. PMID: 40182550 Free PMC article.
-
Comparative analysis of chloroplast genomes on Meliaceae species: insights into the evolution and species identification.Front Plant Sci. 2025 Mar 11;16:1536313. doi: 10.3389/fpls.2025.1536313. eCollection 2025. Front Plant Sci. 2025. PMID: 40134614 Free PMC article.
-
Phylogenetic analysis of nine Impatiens species from subgenus Clavicarpa and subgenus Impatiens (Sect. Impatiens and Sect. Racemosae) based on chloroplast genomes.Front Plant Sci. 2025 Mar 27;16:1541320. doi: 10.3389/fpls.2025.1541320. eCollection 2025. Front Plant Sci. 2025. PMID: 40212867 Free PMC article.
References
-
- Bierhorst D. W. (1958). Vessels in Equisetum . Am. J. Bot. 45, 534–537. doi: 10.2307/2439576 - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

