Nitrate transporter protein NPF5.12 and major latex-like protein MLP6 are important defense factors against Verticillium longisporum
- PMID: 38666306
- PMCID: PMC11233413
- DOI: 10.1093/jxb/erae185
Nitrate transporter protein NPF5.12 and major latex-like protein MLP6 are important defense factors against Verticillium longisporum
Abstract
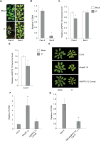
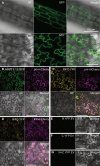
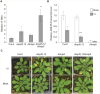
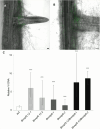
Plant defense responses to the soil-borne fungus Verticillium longisporum causing stem stripe disease on oilseed rape (Brassica napus) are poorly understood. In this study, a population of recombinant inbred lines (RILs) using the Arabidopsis accessions Sei-0 and Can-0 was established. Composite interval mapping, transcriptome data, and T-DNA mutant screening identified the NITRATE/PEPTIDE TRANSPORTER FAMILY 5.12 (AtNPF5.12) gene as being associated with disease susceptibility in Can-0. Co-immunoprecipitation revealed interaction between AtNPF5.12 and the MAJOR LATEX PROTEIN family member AtMLP6, and fluorescence microscopy confirmed this interaction in the plasma membrane and endoplasmic reticulum. CRISPR/Cas9 technology was applied to mutate the NPF5.12 and MLP6 genes in B. napus. Elevated fungal growth in the npf5.12 mlp6 double mutant of both oilseed rape and Arabidopsis demonstrated the importance of these genes in defense against V. longisporum. Colonization of this fungus depends also on available nitrates in the host root. Accordingly, the negative effect of nitrate depletion on fungal growth was less pronounced in Atnpf5.12 plants with impaired nitrate transport. In addition, suberin staining revealed involvement of the NPF5.12 and MLP6 genes in suberin barrier formation. Together, these results demonstrate a dependency on multiple plant factors that leads to successful V. longisporum root infection.
Keywords: Brassica napus; Verticillium longisporum; Arabidopsis; MLP6; NPF5.12; major latex protein; nitrate.
© The Author(s) 2024. Published by Oxford University Press on behalf of the Society for Experimental Biology.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures

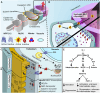
References
-
- Alonso JM, Stepanova AN, Leisse TJ, et al. 2003. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657. - PubMed
-
- Barberon M, Vermeer J, De Bellis D, et al. 2016. Adaptation of root function by nutrient-induced plasticity of endodermal differentiation. Cell 164, 447–459. - PubMed
-
- Benjamini Y, Hochberg Y.. 1995. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society. Series B 57, 289–300.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

