Genomic Epidemiology of C2/H30Rx and C1-M27 Subclades of Escherichia coli ST131 Isolates from Clinical Blood Samples in Hungary
- PMID: 38667039
- PMCID: PMC11047377
- DOI: 10.3390/antibiotics13040363
Genomic Epidemiology of C2/H30Rx and C1-M27 Subclades of Escherichia coli ST131 Isolates from Clinical Blood Samples in Hungary
Abstract
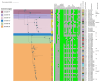
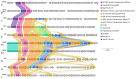
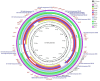
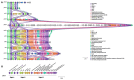
Extended-spectrum β-lactamase-producing Escherichia coli ST131 has become widespread worldwide. This study aims to characterize the virulome, resistome, and population structure of E. coli ST131 isolates from clinical blood samples in Hungary. A total of 30 C2/H30Rx and 33 C1-M27 ST131 isolates were selected for Illumina MiSeq sequencing and 30 isolates for MinION sequencing, followed by hybrid de novo assembly. Five C2/H30Rx and one C1-M27 cluster were identified. C1-M27 isolates harbored the F1:A2:B20 plasmid in 93.9% of cases. Long-read sequencing revealed that blaCTX-M-27 was on plasmids. Among the C2/H30Rx isolates, only six isolates carried the C2-associated F2:A1:B- plasmid type. Of 19 hybrid-assembled C2/H30Rx genomes, the blaCTX-M-15 gene was located on plasmid only in one isolate, while in the other isolates, ISEcp1 or IS26-mediated chromosomal integration of blaCTX-M-15 was detected in unique variations. In one isolate a part of F2:A1:B- plasmid integrated into the chromosome. These results suggest that CTX-M-15-producing C2/H30Rx and CTX-M-27-producing C1-M27 subclades may have emerged and spread in different ways in Hungary. While blaCTX-M-27 was carried mainly on the C1/H30R-associated F1:A2:B20 plasmid, the IncF-like plasmids of C2/H30Rx or its composite transposons have been incorporated into the chromosome through convergent evolutionary processes.
Keywords: C1-M27; C2/H30RX; Escherichia coli; ST131; blaCTX-M-15; blaCTX-M-27; long-read sequencing; whole genome sequencing (WGS).
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
Population snapshot of the extended-spectrum β-lactamase-producing Escherichia coli invasive strains isolated from a Hungarian hospital.Ann Clin Microbiol Antimicrob. 2022 Feb 10;21(1):3. doi: 10.1186/s12941-022-00493-8. Ann Clin Microbiol Antimicrob. 2022. PMID: 35144632 Free PMC article.
-
Short- and Long-Read Sequencing Reveals the Presence and Evolution of an IncF Plasmid Harboring blaCTX-M-15 and blaCTX-M-27 Genes in Escherichia coli ST131.Microbiol Spectr. 2023 Aug 17;11(4):e0035623. doi: 10.1128/spectrum.00356-23. Epub 2023 Jul 19. Microbiol Spectr. 2023. PMID: 37466446 Free PMC article.
-
High Prevalence of ST131 Subclades C2-H30Rx and C1-M27 Among Extended-Spectrum β-Lactamase-Producing Escherichia coli Causing Human Extraintestinal Infections in Patients From Two Hospitals of Spain and France During 2015.Front Cell Infect Microbiol. 2020 Mar 24;10:125. doi: 10.3389/fcimb.2020.00125. eCollection 2020. Front Cell Infect Microbiol. 2020. PMID: 32266173 Free PMC article.
-
Escherichia coli ST131: a multidrug-resistant clone primed for global domination.F1000Res. 2017 Feb 28;6:F1000 Faculty Rev-195. doi: 10.12688/f1000research.10609.1. eCollection 2017. F1000Res. 2017. PMID: 28344773 Free PMC article. Review.
-
Extended-Spectrum β-Lactamase-Producing Enterobacteriaceae: Update on Molecular Epidemiology and Treatment Options.Drugs. 2019 Sep;79(14):1529-1541. doi: 10.1007/s40265-019-01180-3. Drugs. 2019. PMID: 31407238 Review.
Cited by
-
Analysis of molecular mechanisms of delafloxacin resistance in Escherichia coli.Sci Rep. 2024 Nov 2;14(1):26423. doi: 10.1038/s41598-024-78124-9. Sci Rep. 2024. PMID: 39488602 Free PMC article.
-
Extended-Spectrum Beta-Lactamase- and Plasmidic AmpC-Producing Enterobacterales among the Faecal Samples in the Bulgarian Community.Microorganisms. 2024 Aug 28;12(9):1777. doi: 10.3390/microorganisms12091777. Microorganisms. 2024. PMID: 39338452 Free PMC article.
-
Pathogenesis and Immunomodulation of Urinary Tract Infections Caused by Uropathogenic Escherichia coli.Microorganisms. 2025 Mar 26;13(4):745. doi: 10.3390/microorganisms13040745. Microorganisms. 2025. PMID: 40284582 Free PMC article. Review.
-
Comprehensive Assessment of Initial Adaptation of Extended-Spectrum β-Lactamase-Positive ST131 Escherichia coli to Carbapenem Exposure.J Infect Dis. 2025 Apr 15;231(4):e685-e696. doi: 10.1093/infdis/jiae587. J Infect Dis. 2025. PMID: 39602497 Free PMC article.
References
-
- Magiorakos A.-P., Srinivasan A., Carey R.B., Carmeli Y., Falagas M.E., Giske C.G., Harbarth S., Hindler J.F., Kahlmeter G., Olsson-Liljequist B., et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012;18:268–281. doi: 10.1111/j.1469-0691.2011.03570.x. - DOI - PubMed
-
- Cassini A., Högberg L.D., Plachouras D., Quattrocchi A., Hoxha A., Simonsen G.S., Colomb-Cotinat M., Kretzschmar M.E., Devleesschauwer B., Cecchini M., et al. Attributable Deaths and Disability-Adjusted Life-Years Caused by Infections with Antibiotic-Resistant Bacteria in the EU and the European Economic Area in 2015: A Population-Level Modelling Analysis. Lancet Infect. Dis. 2019;19:56–66. doi: 10.1016/S1473-3099(18)30605-4. - DOI - PMC - PubMed
-
- Petty N.K., Ben Zakour N.L., Stanton-Cook M., Skippington E., Totsika M., Forde B.M., Phan M.-D., Gomes Moriel D., Peters K.M., Davies M., et al. Global Dissemination of a Multidrug Resistant Escherichia coli Clone. Proc. Natl. Acad. Sci. USA. 2014;111:5694–5699. doi: 10.1073/pnas.1322678111. - DOI - PMC - PubMed
-
- Johnson T.J., Danzeisen J.L., Youmans B., Case K., Llop K., Munoz-Aguayo J., Flores-Figueroa C., Aziz M., Stoesser N., Sokurenko E., et al. Separate F-Type Plasmids Have Shaped the Evolution of the H30 Subclone of Escherichia coli Sequence Type 131. mSphere. 2016;1:e00121-16. doi: 10.1128/mSphere.00121-16. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

