Regulation of Mertk Surface Expression via ADAM17 and γ-Secretase Proteolytic Processing
- PMID: 38673989
- PMCID: PMC11050108
- DOI: 10.3390/ijms25084404
Regulation of Mertk Surface Expression via ADAM17 and γ-Secretase Proteolytic Processing
Abstract
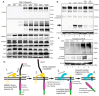
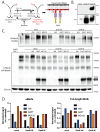
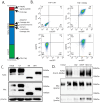
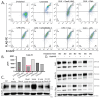
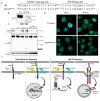
Mertk, a type I receptor tyrosine kinase and member of the TAM family of receptors, has important functions in promoting efferocytosis and resolving inflammation under physiological conditions. In recent years, Mertk has also been linked to pathophysiological roles in cancer, whereby, in several cancer types, including solid cancers and leukemia/lymphomas. Mertk contributes to oncogenic features of proliferation and cell survival as an oncogenic tyrosine kinase. In addition, Mertk expressed on macrophages, including tumor-associated macrophages, promotes immune evasion in cancer and is suggested to act akin to a myeloid checkpoint inhibitor that skews macrophages towards inhibitory phenotypes that suppress host T-cell anti-tumor immunity. In the present study, to better understand the post-translational regulation mechanisms controlling Mertk expression in monocytes/macrophages, we used a PMA-differentiated THP-1 cell model to interrogate the regulation of Mertk expression and developed a novel Mertk reporter cell line to study the intracellular trafficking of Mertk. We show that PMA treatment potently up-regulates Mertk as well as components of the ectodomain proteolytic processing platform ADAM17, whereas PMA differentially regulates the canonical Mertk ligands Gas6 and Pros1 (Gas6 is down-regulated and Pros1 is up-regulated). Under non-stimulated homeostatic conditions, Mertk in PMA-differentiated THP1 cells shows active constitutive proteolytic cleavage by the sequential activities of ADAM17 and the Presenilin/γ-secretase complex, indicating that Mertk is cleaved homeostatically by the combined sequential action of ADAM17 and γ-secretase, after which the cleaved intracellular fragment of Mertk is degraded in a proteasome-dependent mechanism. Using chimeric Flag-Mertk-EGFP-Myc reporter receptors, we confirm that inhibitors of γ-secretase and MG132, which inhibits the 26S proteasome, stabilize the intracellular fragment of Mertk without evidence of nuclear translocation. Finally, the treatment of cells with active γ-carboxylated Gas6, but not inactive Warfarin-treated non-γ-carboxylated Gas6, regulates a distinct proteolytic itinerary-involved receptor clearance and lysosomal proteolysis. Together, these results indicate that pleotropic and complex proteolytic activities regulate Mertk ectodomain cleavage as a homeostatic negative regulatory event to safeguard against the overactivation of Mertk.
Keywords: ADAM-17 ectodomain shedding; Gas6; Mertk; intracellular receptor trafficking; membrane expression; proteolysis; γ-secretase.
Conflict of interest statement
The authors declare no conflicts of interest. The funders had no role in the design of the study, in the collection, analyses, or interpretation of data, in the writing of the manuscript, or in the decision to publish the results.
Figures


Similar articles
-
Expression of TAM-R in Human Immune Cells and Unique Regulatory Function of MerTK in IL-10 Production by Tolerogenic DC.Front Immunol. 2020 Sep 25;11:564133. doi: 10.3389/fimmu.2020.564133. eCollection 2020. Front Immunol. 2020. PMID: 33101282 Free PMC article.
-
A Disintegrin and Metalloproteinase 17 Disrupts Bovine Macrophage MER Proto-Oncogene Tyrosine Kinase Integrity to Impede Apoptotic Cell Clearance and Promote Inflammation in Clinical Mastitis.J Agric Food Chem. 2025 Jan 8;73(1):549-561. doi: 10.1021/acs.jafc.4c09164. Epub 2024 Dec 28. J Agric Food Chem. 2025. PMID: 39731564
-
Shedding of the Mer tyrosine kinase receptor is mediated by ADAM17 protein through a pathway involving reactive oxygen species, protein kinase Cδ, and p38 mitogen-activated protein kinase (MAPK).J Biol Chem. 2011 Sep 23;286(38):33335-44. doi: 10.1074/jbc.M111.263020. Epub 2011 Aug 2. J Biol Chem. 2011. PMID: 21828049 Free PMC article.
-
Post-translational modifications of the ligands: Requirement for TAM receptor activation.Int Rev Cell Mol Biol. 2020;357:35-55. doi: 10.1016/bs.ircmb.2020.09.002. Epub 2020 Oct 27. Int Rev Cell Mol Biol. 2020. PMID: 33234244 Review.
-
Targeting Tyro3, Axl and MerTK (TAM receptors): implications for macrophages in the tumor microenvironment.Mol Cancer. 2019 May 14;18(1):94. doi: 10.1186/s12943-019-1022-2. Mol Cancer. 2019. PMID: 31088471 Free PMC article. Review.
Cited by
-
Functions of TAM Receptors and Ligands Protein S and Gas6 in Atherosclerosis and Cardiovascular Disease.Int J Mol Sci. 2024 Nov 27;25(23):12736. doi: 10.3390/ijms252312736. Int J Mol Sci. 2024. PMID: 39684449 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

