Repeated Occurrence of Mobile Colistin Resistance Gene-Carrying Plasmids in Pathogenic Escherichia coli from German Pig Farms
- PMID: 38674671
- PMCID: PMC11052496
- DOI: 10.3390/microorganisms12040729
Repeated Occurrence of Mobile Colistin Resistance Gene-Carrying Plasmids in Pathogenic Escherichia coli from German Pig Farms
Abstract
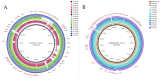
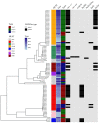
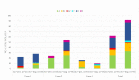
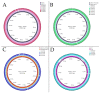
The global spread of plasmid-mediated mobile colistin resistance (mcr) genes threatens the vital role of colistin as a drug of last resort. We investigated whether the recurrent occurrence of specific E. coli pathotypes and plasmids in individual pig farms resulted from the continued presence or repeated reintroduction of distinct E. coli strains. E. coli isolates (n = 154) obtained from three pig farms with at least four consecutive years of mcr detection positive for virulence-associated genes (VAGs) predicting an intestinal pathogenic pathotype via polymerase chain reaction were analyzed. Detailed investigation of VAGs, antimicrobial resistance genes and plasmid Inc types was conducted using whole genome sequencing for 87 selected isolates. Sixty-one E. coli isolates harbored mcr-1, and one isolate carried mcr-4. On Farm 1, mcr-positive isolates were either edema disease E. coli (EDEC; 77.3%) or enterotoxigenic E. coli (ETEC; 22.7%). On Farm 2, all mcr-positive strains were ETEC, while mcr-positive isolates from Farm 3 showed a wider range of pathotypes. The mcr-1.1 gene was located on IncHI2 (Farm 1), IncX4 (Farm 2) or IncX4 and IncI2 plasmids (Farm 3). These findings suggest that various pathogenic E. coli strains play an important role in maintaining plasmid-encoded colistin resistance genes in the pig environment over time.
Keywords: EDEC; ETEC; Escherichia coli; IncHI2; IncI2; IncX4; mcr-1; mobile colistin resistance; plasmid; swine.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
References
-
- García-Meniño I., García V., Mora A., Díaz-Jiménez D., Flament-Simon S.C., Alonso M.P., Blanco J.E., Blanco M., Blanco J. Swine Enteric Colibacillosis in Spain: Pathogenic Potential of mcr-1 ST10 and ST131 E. coli Isolates. Front. Microbiol. 2018;9:2659. doi: 10.3389/fmicb.2018.02659. - DOI - PMC - PubMed
-
- Fairbrother J.M., Gyles C.L. Colibacillosis. In: Zimmerman J.J., Karriker L.A., Ramirez A., Schwartz K.J., Stevenson G.W., editors. Disease of Swine. 10th ed. Wiley; Hoboken, NJ, USA: 2012. pp. 723–747.
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

