Current Perspectives on Biological Screening of Newly Synthetised Sulfanilamide Schiff Bases as Promising Antibacterial and Antibiofilm Agents
- PMID: 38675368
- PMCID: PMC11053482
- DOI: 10.3390/ph17040405
Current Perspectives on Biological Screening of Newly Synthetised Sulfanilamide Schiff Bases as Promising Antibacterial and Antibiofilm Agents
Abstract
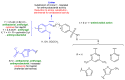
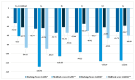
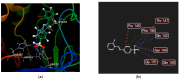
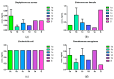
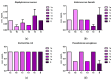
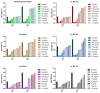
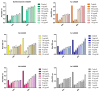
Growing resistance to antimicrobials, combined with pathogens that form biofilms, presents significant challenges in healthcare. Modifying current antimicrobial agents is an economical approach to developing novel molecules that could exhibit biological activity. Thus, five sulfanilamide Schiff bases were synthesized under microwave irradiation and characterized spectroscopically and in silico. They were evaluated for their antimicrobial and antibiofilm activities against both Gram-positive and Gram-negative bacterial strains. Their cytotoxic potential against two cancer cell lines was also determined. Gram-positive bacteria were susceptible to the action of these compounds. Derivatives 1b and 1d inhibited S. aureus's growth (MIC from 0.014 mg/mL) and biofilm (IC from 0.029 mg/mL), while compound 1e was active against E. faecalis's planktonic and sessile forms. Two compounds significantly reduced cell viability at 5 μg/mL after 24 h of exposure (1d-HT-29 colorectal adenocarcinoma cells, 1c-LN229 glioblastoma cells). A docking study revealed the increased binding affinities of these derivatives compared to sulfanilamide. Hence, these Schiff bases exhibited higher activity compared to their parent drug, with halogen groups playing a crucial role in both their antimicrobial and cytotoxic effects.
Keywords: Schiff base; antibiofilm; antimicrobial; cytotoxicity; sulfanilamide.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

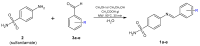



Similar articles
-
In vitro and in vivo evaluation of the antimicrobial, antioxidant, cytotoxic, hemolytic activities and in silico POM/DFT/DNA-binding and pharmacokinetic analyses of new sulfonamide bearing thiazolidin-4-ones.J Biomol Struct Dyn. 2024 Apr;42(7):3747-3763. doi: 10.1080/07391102.2023.2226713. Epub 2023 Jul 4. J Biomol Struct Dyn. 2024. PMID: 37402503
-
Antimicrobial and cytotoxic activities of isoniazid connected menthone derivatives and their investigation of clinical pathogens causing infectious disease.J Infect Public Health. 2021 Apr;14(4):533-542. doi: 10.1016/j.jiph.2020.12.033. Epub 2021 Feb 6. J Infect Public Health. 2021. PMID: 33744741
-
Benzimidazole Schiff base derivatives: synthesis, characterization and antimicrobial activity.BMC Chem. 2019 Nov 9;13(1):127. doi: 10.1186/s13065-019-0642-3. eCollection 2019 Dec. BMC Chem. 2019. PMID: 31728454 Free PMC article.
-
Antimicrobial and antibiofilm activities of Cu(II) Schiff base complexes against methicillin-susceptible and resistant Staphylococcus aureus.Ann Clin Microbiol Antimicrob. 2021 Sep 24;20(1):67. doi: 10.1186/s12941-021-00473-4. Ann Clin Microbiol Antimicrob. 2021. PMID: 34560892 Free PMC article.
-
Small Schiff Base Molecules-A Possible Strategy to Combat Biofilm-Related Infections.Antibiotics (Basel). 2024 Jan 12;13(1):75. doi: 10.3390/antibiotics13010075. Antibiotics (Basel). 2024. PMID: 38247634 Free PMC article. Review.
References
-
- New WHO Report Highlights Progress, but Also Remaining Gaps, in Ensuring a Robust Pipeline of Antibiotic Treatments to Combat Antimicrobial Resistance (AMR) [(accessed on 23 May 2023)]. Available online: https://www.who.int/news/item/15-05-2023-new-who-report-highlights-progr...
-
- Høiby N., Bjarnsholt T., Moser C., Bassi G.L., Coenye T., Donelli G., Hall-Stoodley L., Holá V., Imbert C., Kirketerp-Møller K., et al. ESCMID* Guideline for the Diagnosis and Treatment of Biofilm Infections 2014. Clin. Microbiol. Infect. 2015;21:S1–S25. doi: 10.1016/j.cmi.2014.10.024. - DOI - PubMed
-
- Ceri H., Gorman S.P., Gilmore B.F. Microbial Biofilms: Consequences for Health. In: Denyer S.P., Hodges N., Gorman S.P., Gilmore B.F., editors. Hugo and Russell’s Pharmaceutical Microbiology. Wiley-Blackwell; Hoboken, NJ, USA: 2011. pp. 121–130.
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

