Investigating Potential Cancer Therapeutics: Insight into Histone Deacetylases (HDACs) Inhibitions
- PMID: 38675404
- PMCID: PMC11054547
- DOI: 10.3390/ph17040444
Investigating Potential Cancer Therapeutics: Insight into Histone Deacetylases (HDACs) Inhibitions
Abstract
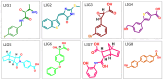
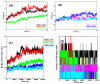
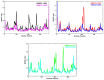
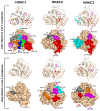
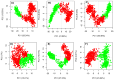
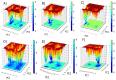
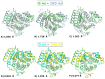
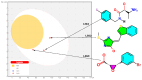
Histone deacetylases (HDACs) are enzymes that remove acetyl groups from ɛ-amino of histone, and their involvement in the development and progression of cancer disorders makes them an interesting therapeutic target. This study seeks to discover new inhibitors that selectively inhibit HDAC enzymes which are linked to deadly disorders like T-cell lymphoma, childhood neuroblastoma, and colon cancer. MOE was used to dock libraries of ZINC database molecules within the catalytic active pocket of target HDACs. The top three hits were submitted to MD simulations ranked on binding affinities and well-occupied interaction mechanisms determined from molecular docking studies. Inside the catalytic active site of HDACs, the two stable inhibitors LIG1 and LIG2 affect the protein flexibility, as evidenced by RMSD, RMSF, Rg, and PCA. MD simulations of HDACs complexes revealed an alteration from extended to bent motional changes within loop regions. The structural deviation following superimposition shows flexibility via a visual inspection of movable loops at different timeframes. According to PCA, the activity of HDACs inhibitors induces structural dynamics that might potentially be utilized to define the nature of protein inhibition. The findings suggest that this study offers solid proof to investigate LIG1 and LIG2 as potential HDAC inhibitors.
Keywords: histone deacetylases; molecular docking; molecular dynamic simulation; neuroblastoma.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
Insights into structural features of HDAC1 and its selectivity inhibition elucidated by Molecular dynamic simulation and Molecular Docking.J Biomol Struct Dyn. 2019 Feb;37(3):584-610. doi: 10.1080/07391102.2018.1441072. Epub 2018 Mar 6. J Biomol Struct Dyn. 2019. PMID: 29447615
-
Phenolic compounds as histone deacetylase inhibitors: binding propensity and interaction insights from molecular docking and dynamics simulations.Amino Acids. 2023 May;55(5):579-593. doi: 10.1007/s00726-023-03249-6. Epub 2023 Feb 12. Amino Acids. 2023. PMID: 36781452
-
Targeting protein tyrosine phosphatase to unravel possible inhibitors for Streptococcus pneumoniae using molecular docking, molecular dynamics simulations coupled with free energy calculations.Life Sci. 2021 Jan 1;264:118621. doi: 10.1016/j.lfs.2020.118621. Epub 2020 Oct 22. Life Sci. 2021. PMID: 33164832
-
Medicinal chemistry advances in targeting class I histone deacetylases.Explor Target Antitumor Ther. 2023;4(4):757-779. doi: 10.37349/etat.2023.00166. Epub 2023 Aug 31. Explor Target Antitumor Ther. 2023. PMID: 37711592 Free PMC article. Review.
-
Histone Deacetylases and Histone Deacetylase Inhibitors in Neuroblastoma.Front Cell Dev Biol. 2020 Oct 7;8:578770. doi: 10.3389/fcell.2020.578770. eCollection 2020. Front Cell Dev Biol. 2020. PMID: 33117806 Free PMC article. Review.
Cited by
-
Decoding the Epigenome of Breast Cancer.Int J Mol Sci. 2025 Mar 13;26(6):2605. doi: 10.3390/ijms26062605. Int J Mol Sci. 2025. PMID: 40141248 Free PMC article. Review.
References
-
- Dawood M., Elbadawi M., Böckers M., Bringmann G., Efferth T. Molecular docking-based virtual drug screening revealing an oxofluorenyl benzamide and a bromonaphthalene sulfonamido hydroxybenzoic acid as HDAC6 inhibitors with cytotoxicity against leukemia cells. Biomed. Pharmacother. 2020;129:110454. doi: 10.1016/j.biopha.2020.110454. - DOI - PubMed
-
- Uba A.I., Hryb M., Singh M., Bui-Linh C., Tran A., Atienza J., Misbah S., Mou X., Wu C. Discovery of novel inhibitors of histone deacetylase 6: Structure-based virtual screening, molecular dynamics simulation, enzyme inhibition and cell viability assays. Life Sci. 2024;338:122395. doi: 10.1016/j.lfs.2023.122395. - DOI - PubMed
-
- Santos-Barriopedro I., Raurell-Vila H., Vaquero A. The role of hats and hdacs in cell physiology and disease. Gene Regul. Epigenetics Horm. Signal. 2017;17:101–136.
-
- Carta F. Metalloenzymes. Academic Press; Cambridge, MA, USA: 2024. Histone deacetylases and other epigenetic targets; pp. 265–281.
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

