Measurable residual disease monitoring by ddPCR in the early posttransplant period complements the traditional MFC method to predict relapse after HSCT in AML/MDS: a multicenter retrospective study
- PMID: 38689269
- PMCID: PMC11061929
- DOI: 10.1186/s12967-024-05114-w
Measurable residual disease monitoring by ddPCR in the early posttransplant period complements the traditional MFC method to predict relapse after HSCT in AML/MDS: a multicenter retrospective study
Abstract
Background: Droplet digital PCR (ddPCR) is widely applied to monitor measurable residual disease (MRD). However, there are limited studies on the feasibility of ddPCR-MRD monitoring after allogeneic hematopoietic stem cell transplantation (allo-HSCT), especially targeting multiple molecular markers simultaneously.
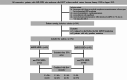
Methods: Our study collected samples from patients with acute myeloid leukemia (AML) or high-risk myelodysplastic syndrome (MDS) in complete remission after allo-HSCT between January 2018 and August 2021 to evaluate whether posttransplant ddPCR-MRD monitoring can identify patients at high risk of relapse.
Results: Of 152 patients, 58 (38.2%) were MRD positive by ddPCR within 4 months posttransplant, with a median variant allele frequency of 0.198%. The detectable DTA mutations (DNMT3A, TET2, and ASXL1 mutations) after allo-HSCT were not associated with an increased risk of relapse. After excluding DTA mutations, patients with ddPCR-MRD positivity had a significantly higher cumulative incidence of relapse (CIR, 38.7% vs. 9.7%, P < 0.001) and lower rates of relapse-free survival (RFS, 55.5% vs. 83.7%, P < 0.001) and overall survival (OS, 60.5% vs. 90.5%, P < 0.001). In multivariate analysis, ddPCR-MRD positivity of non-DTA genes was an independent adverse predictor for CIR (hazard ratio [HR], 4.02; P < 0.001), RFS (HR, 2.92; P = 0.002) and OS (HR, 3.12; P = 0.007). Moreover, the combination of ddPCR with multiparameter flow cytometry (MFC) can further accurately identify patients at high risk of relapse (F+/M+, HR, 22.44; P < 0.001, F+/M-, HR, 12.46; P < 0.001 and F-/M+, HR, 4.51; P = 0.003).
Conclusion: ddPCR-MRD is a feasible approach to predict relapse after allo-HSCT in AML/MDS patients with non-DTA genes and is more accurate when combined with MFC.
Trial registration: ClinicalTrials.gov identifier: NCT06000306. Registered 17 August 2023 -Retrospectively registered ( https://clinicaltrials.gov/study/NCT06000306?term=NCT06000306&rank=1 ).
Keywords: Allo-HSCT; MFC; MRD; Relapse; ddPCR.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Patient-Specific Measurable Residual Disease Markers Predict Outcome in Patients With Myelodysplastic Syndrome and Related Diseases After Hematopoietic Stem-Cell Transplantation.J Clin Oncol. 2024 Apr 20;42(12):1378-1390. doi: 10.1200/JCO.23.01159. Epub 2024 Jan 17. J Clin Oncol. 2024. PMID: 38232336
-
Monitoring of post-transplant MLL-PTD as minimal residual disease can predict relapse after allogeneic HSCT in patients with acute myeloid leukemia and myelodysplastic syndrome.BMC Cancer. 2022 Jan 3;22(1):11. doi: 10.1186/s12885-021-09051-5. BMC Cancer. 2022. PMID: 34979982 Free PMC article.
-
[Effect of minimal residual disease monitoring by multiparameter flow cytometry pre-conditioning on prognosis of acute myeloid leukemia after allogeneic hematopoietic stem cell transplantation].Zhonghua Xue Ye Xue Za Zhi. 2017 Feb 14;38(2):118-123. doi: 10.3760/cma.j.issn.0253-2727.2017.02.007. Zhonghua Xue Ye Xue Za Zhi. 2017. PMID: 28279035 Free PMC article. Chinese.
-
Test Then Erase? Current Status and Future Opportunities for Measurable Residual Disease Testing in Acute Myeloid Leukemia.Acta Haematol. 2024;147(2):133-146. doi: 10.1159/000535463. Epub 2023 Nov 30. Acta Haematol. 2024. PMID: 38035547 Free PMC article. Review.
-
Pre-emptive therapeutic decisions based on measurable residual disease status in acute myeloid leukemia: ready for prime time?Leukemia. 2025 Jan;39(1):1-7. doi: 10.1038/s41375-024-02458-6. Epub 2024 Nov 5. Leukemia. 2025. PMID: 39496917 Review.
Cited by
-
Digital PCR: from early developments to its future application in clinics.Lab Chip. 2025 Aug 5;25(16):3921-3961. doi: 10.1039/d5lc00055f. Lab Chip. 2025. PMID: 40686367 Free PMC article. Review.
-
Combination of disease burden before allogeneic transplantation and early post-transplant minimal residual disease predicts survival in patients with acute myeloid leukemia.Ann Hematol. 2025 Apr;104(4):2469-2481. doi: 10.1007/s00277-025-06325-x. Epub 2025 Apr 25. Ann Hematol. 2025. PMID: 40278922 Free PMC article.
References
-
- Platzbecker U, Middeke JM, Sockel K, Herbst R, Wolf D, Baldus CD, et al. Measurable residual disease-guided treatment with azacitidine to prevent haematological relapse in patients with myelodysplastic syndrome and acute myeloid leukaemia (RELAZA2): an open-label, multicentre, phase 2 trial. Lancet Oncol. 2018;19(12):1668–79. doi: 10.1016/S1470-2045(18)30580-1. - DOI - PubMed
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

