Investigating the genetic control of plant development in spring barley under speed breeding conditions
- PMID: 38691245
- PMCID: PMC11063105
- DOI: 10.1007/s00122-024-04618-9
Investigating the genetic control of plant development in spring barley under speed breeding conditions
Abstract
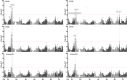
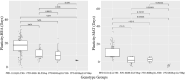
This study found that the genes, PPD-H1 and ELF3, control the acceleration of plant development under speed breeding, with important implications for optimizing the delivery of climate-resilient crops. Speed breeding is a tool to accelerate breeding and research programmes. Despite its success and growing popularity with breeders, the genetic basis of plant development under speed breeding remains unknown. This study explored the developmental advancements of barley genotypes under different photoperiod regimes. A subset of the HEB-25 Nested Association Mapping population was evaluated for days to heading and maturity under two contrasting photoperiod conditions: (1) Speed breeding (SB) consisting of 22 h of light and 2 h of darkness, and (2) normal breeding (NB) consisting of 16 h of light and 8 h of darkness. GWAS revealed that developmental responses under both conditions were largely controlled by two loci: PPDH-1 and ELF3. Allelic variants at these genes determine whether plants display early flowering and maturity under both conditions. At key QTL regions, domesticated alleles were associated with late flowering and maturity in NB and early flowering and maturity in SB, whereas wild alleles were associated with early flowering under both conditions. We hypothesize that this is related to the dark-dependent repression of PPD-H1 by ELF3 which might be more prominent in NB conditions. Furthermore, by comparing development under two photoperiod regimes, we derived an estimate of plasticity for the two traits. Interestingly, plasticity in development was largely attributed to allelic variation at ELF3. Our results have important implications for our understanding and optimization of speed breeding protocols particularly for introgression breeding and the design of breeding programmes to support the delivery of climate-resilient crops.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no confict of interest.
Figures

Similar articles
-
Genetic dissection of photoperiod response based on GWAS of pre-anthesis phase duration in spring barley.PLoS One. 2014 Nov 24;9(11):e113120. doi: 10.1371/journal.pone.0113120. eCollection 2014. PLoS One. 2014. PMID: 25420105 Free PMC article.
-
Association of barley photoperiod and vernalization genes with QTLs for flowering time and agronomic traits in a BC2DH population and a set of wild barley introgression lines.Theor Appl Genet. 2010 May;120(8):1559-74. doi: 10.1007/s00122-010-1276-y. Epub 2010 Feb 13. Theor Appl Genet. 2010. PMID: 20155245 Free PMC article.
-
Diverse alleles of Photoperiod-H1 directly and indirectly affect barley yield-related traits under contrasting photoperiods and PHYTOCHROME C backgrounds.J Exp Bot. 2025 Apr 9;76(6):1678-1690. doi: 10.1093/jxb/erae491. J Exp Bot. 2025. PMID: 39851238
-
Major flowering time genes of barley: allelic diversity, effects, and comparison with wheat.Theor Appl Genet. 2021 Jul;134(7):1867-1897. doi: 10.1007/s00122-021-03824-z. Epub 2021 May 9. Theor Appl Genet. 2021. PMID: 33969431 Free PMC article. Review.
-
Genome-wide association study of heading and flowering dates and construction of its prediction equation in Chinese common wheat.Theor Appl Genet. 2018 Nov;131(11):2271-2285. doi: 10.1007/s00122-018-3181-8. Epub 2018 Sep 14. Theor Appl Genet. 2018. PMID: 30218294 Review.
Cited by
-
Integrating molecular genetics with plant breeding to deliver impact.Plant Physiol. 2025 Jul 3;198(3):kiaf087. doi: 10.1093/plphys/kiaf087. Plant Physiol. 2025. PMID: 40036799 Free PMC article. Review.
-
Speed-bred crops for food security and sustainable agriculture.Planta. 2025 Jun 19;262(2):34. doi: 10.1007/s00425-025-04746-6. Planta. 2025. PMID: 40536536 Review.
References
-
- Arthur JM, Guthrie JD, Newell JM. Some effects of artificial climates on the growth and chemical composition of plants. Am J Bot. 1930;17(5):416. doi: 10.2307/2435930. - DOI
-
- Bates D, Mächler M, Bolker B, Walker S. Fitting linear mixed-effects models using lme4. J Stat Softw. 2015 doi: 10.18637/jss.v067.i01. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

