Nucleic Acid-Binding Dyes as Versatile Photocatalysts for Atom-Transfer Radical Polymerization
- PMID: 38691811
- PMCID: PMC11100002
- DOI: 10.1021/jacs.4c03513
Nucleic Acid-Binding Dyes as Versatile Photocatalysts for Atom-Transfer Radical Polymerization
Abstract
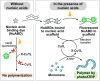
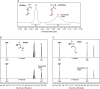
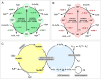
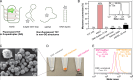
Nucleic acid-binding dyes (NuABDs) are fluorogenic probes that light up after binding to nucleic acids. Taking advantage of their fluorogenicity, NuABDs have been widely utilized in the fields of nanotechnology and biotechnology for diagnostic and analytical applications. We demonstrate the potential of NuABDs together with an appropriate nucleic acid scaffold as an intriguing photocatalyst for precisely controlled atom-transfer radical polymerization (ATRP). Additionally, we systematically investigated the thermodynamic and electrochemical properties of the dyes, providing insights into the mechanism that drives the photopolymerization. The versatility of the NuABD-based platform was also demonstrated through successful polymerizations using several NuABDs in conjunction with diverse nucleic acid scaffolds, such as G-quadruplex DNA or DNA nanoflowers. This study not only extends the horizons of controlled photopolymerization but also broadens opportunities for nucleic acid-based materials and technologies, including nucleic acid-polymer biohybrids and stimuli-responsive ATRP platforms.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

Similar articles
-
Atom Transfer Radical Polymerization (ATRP) Catalyzed by Visible Light-Absorbed Small Molecule Organic Semiconductors.Macromol Rapid Commun. 2018 Sep;39(18):e1800466. doi: 10.1002/marc.201800466. Epub 2018 Jul 30. Macromol Rapid Commun. 2018. PMID: 30062811
-
Rationally Designed G-Quadruplex Selective "Turn-On" NIR Fluorescent Probe with Large Stokes Shift for Nucleic Acid Research-Based Applications.ACS Appl Bio Mater. 2024 Nov 18;7(11):7233-7243. doi: 10.1021/acsabm.4c00940. Epub 2024 Oct 28. ACS Appl Bio Mater. 2024. PMID: 39466599
-
Visible Light Photochemical Reactions for Nucleic Acid-Based Technologies.Molecules. 2021 Jan 21;26(3):556. doi: 10.3390/molecules26030556. Molecules. 2021. PMID: 33494512 Free PMC article. Review.
-
Antimony-Based Halide Perovskite Nanoparticles as Lead-Free Photocatalysts for Controlled Radical Polymerization.Macromol Rapid Commun. 2024 May;45(9):e2300695. doi: 10.1002/marc.202300695. Epub 2024 Feb 17. Macromol Rapid Commun. 2024. PMID: 38350418
-
Atom Transfer Radical Polymerization: Billion Times More Active Catalysts and New Initiation Systems.Macromol Rapid Commun. 2019 Jan;40(1):e1800616. doi: 10.1002/marc.201800616. Epub 2018 Oct 30. Macromol Rapid Commun. 2019. PMID: 30375120 Review.
Cited by
-
Highly efficient dual photoredox/copper catalyzed atom transfer radical polymerization achieved through mechanism-driven photocatalyst design.Nat Commun. 2024 Jun 17;15(1):5160. doi: 10.1038/s41467-024-49509-1. Nat Commun. 2024. PMID: 38886349 Free PMC article.
References
-
- Gomollón-Bel F. Ten Chemical Innovations That Will Change Our World: IUPAC identifies emerging technologies in Chemistry with potential to make our planet more sustainable. Chem. Int. 2019, 41 (2), 12–17. 10.1515/ci-2019-0203. - DOI
-
- Corrigan N.; Jung K.; Moad G.; Hawker C. J.; Matyjaszewski K.; Boyer C. Reversible-deactivation radical polymerization (Controlled/living radical polymerization): From discovery to materials design and applications. Prog. Polym. Sci. 2020, 111, 101311.10.1016/j.progpolymsci.2020.101311. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

