Prioritizing genomic variants through neuro-symbolic, knowledge-enhanced learning
- PMID: 38696757
- PMCID: PMC11132820
- DOI: 10.1093/bioinformatics/btae301
Prioritizing genomic variants through neuro-symbolic, knowledge-enhanced learning
Abstract
Motivation: Whole-exome and genome sequencing have become common tools in diagnosing patients with rare diseases. Despite their success, this approach leaves many patients undiagnosed. A common argument is that more disease variants still await discovery, or the novelty of disease phenotypes results from a combination of variants in multiple disease-related genes. Interpreting the phenotypic consequences of genomic variants relies on information about gene functions, gene expression, physiology, and other genomic features. Phenotype-based methods to identify variants involved in genetic diseases combine molecular features with prior knowledge about the phenotypic consequences of altering gene functions. While phenotype-based methods have been successfully applied to prioritizing variants, such methods are based on known gene-disease or gene-phenotype associations as training data and are applicable to genes that have phenotypes associated, thereby limiting their scope. In addition, phenotypes are not assigned uniformly by different clinicians, and phenotype-based methods need to account for this variability.
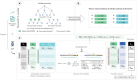
Results: We developed an Embedding-based Phenotype Variant Predictor (EmbedPVP), a computational method to prioritize variants involved in genetic diseases by combining genomic information and clinical phenotypes. EmbedPVP leverages a large amount of background knowledge from human and model organisms about molecular mechanisms through which abnormal phenotypes may arise. Specifically, EmbedPVP incorporates phenotypes linked to genes, functions of gene products, and the anatomical site of gene expression, and systematically relates them to their phenotypic effects through neuro-symbolic, knowledge-enhanced machine learning. We demonstrate EmbedPVP's efficacy on a large set of synthetic genomes and genomes matched with clinical information.
Availability and implementation: EmbedPVP and all evaluation experiments are freely available at https://github.com/bio-ontology-research-group/EmbedPVP.
© The Author(s) 2024. Published by Oxford University Press.
Conflict of interest statement
The authors declare that no conflicts of interest exist.
Figures
Similar articles
-
DeepSVP: integration of genotype and phenotype for structural variant prioritization using deep learning.Bioinformatics. 2022 Mar 4;38(6):1677-1684. doi: 10.1093/bioinformatics/btab859. Bioinformatics. 2022. PMID: 34951628 Free PMC article.
-
A clinical knowledge graph-based framework to prioritize candidate genes for facilitating diagnosis of Mendelian diseases and rare genetic conditions.BMC Bioinformatics. 2025 Mar 14;26(1):82. doi: 10.1186/s12859-025-06096-2. BMC Bioinformatics. 2025. PMID: 40087567 Free PMC article.
-
Predicting candidate genes from phenotypes, functions and anatomical site of expression.Bioinformatics. 2021 May 5;37(6):853-860. doi: 10.1093/bioinformatics/btaa879. Bioinformatics. 2021. PMID: 33051643 Free PMC article.
-
In Silico Functional Annotation of Genomic Variation.Curr Protoc Hum Genet. 2016 Jan 1;88:6.15.1-6.15.17. doi: 10.1002/0471142905.hg0615s88. Curr Protoc Hum Genet. 2016. PMID: 26724722 Free PMC article. Review.
-
Disease insights through cross-species phenotype comparisons.Mamm Genome. 2015 Oct;26(9-10):548-55. doi: 10.1007/s00335-015-9577-8. Epub 2015 Jun 20. Mamm Genome. 2015. PMID: 26092691 Free PMC article. Review.
Cited by
-
GeOKG: geometry-aware knowledge graph embedding for Gene Ontology and genes.Bioinformatics. 2025 Mar 29;41(4):btaf160. doi: 10.1093/bioinformatics/btaf160. Bioinformatics. 2025. PMID: 40217132 Free PMC article.
-
Computational strategies for cross-species knowledge transfer and translational biomedicine.ArXiv [Preprint]. 2024 Aug 16:arXiv:2408.08503v1. ArXiv. 2024. PMID: 39184546 Free PMC article. Preprint.
-
The Unified Phenotype Ontology (uPheno): A framework for cross-species integrative phenomics.bioRxiv [Preprint]. 2024 Sep 22:2024.09.18.613276. doi: 10.1101/2024.09.18.613276. bioRxiv. 2024. Update in: Genetics. 2025 Mar 17;229(3):iyaf027. doi: 10.1093/genetics/iyaf027. PMID: 39345458 Free PMC article. Updated. Preprint.
-
The Unified Phenotype Ontology : a framework for cross-species integrative phenomics.Genetics. 2025 Mar 17;229(3):iyaf027. doi: 10.1093/genetics/iyaf027. Genetics. 2025. PMID: 40048704 Free PMC article.
References
-
- Ali M, Berrendorf M, Hoyt CT. et al. Bringing light into the dark: a large-scale evaluation of knowledge graph embedding models under a unified framework. IEEE Trans Pattern Anal Mach Intell 2021a;44:8825–45. - PubMed
-
- Ali M, Berrendorf M, Hoyt CT. et al. PyKEEN 1.0: a Python library for training and evaluating knowledge graph embeddings. J Mach Learn Res 2021b;22:3723–8.
-
- Amberger J, Bocchini C, Hamosh A. et al. A new face and new challenges for Online Mendelian Inheritance in Man (OMIM®). Hum Mutat 2011;32:564–7. - PubMed

