Individual disruption of 12 testis-enriched genes via the CRISPR/Cas9 system does not affect the fertility of male mice
- PMID: 38697008
- PMCID: PMC11390273
- DOI: 10.1016/j.jri.2024.104252
Individual disruption of 12 testis-enriched genes via the CRISPR/Cas9 system does not affect the fertility of male mice
Abstract
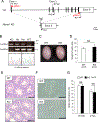
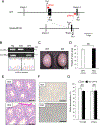
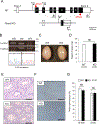
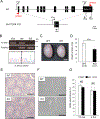
More than 1200 genes have been shown in the database to be expressed predominantly in the mouse testes. Advances in genome editing technologies such as the CRISPR/Cas9 system have made it possible to create genetically engineered mice more rapidly and efficiently than with conventional methods, which can be utilized to screen genes essential for male fertility by knocking out testis-enriched genes. Finding such genes related to male fertility would not only help us understand the etiology of human infertility but also lead to the development of male contraceptives. In this study, we generated knockout mice for 12 genes (Acrv1, Adgrf3, Atp8b5, Cfap90, Cfap276, Fbxw5, Gm17266, Lrrd1, Mroh7, Nemp1, Spata45, and Trim36) that are expressed predominantly in the testis and examined the appearance and histological morphology of testes, sperm motility, and male fertility. Mating tests revealed that none of these genes is essential for male fertility at least individually. Notably, knockout mice for Gm17266 showed smaller testis size than the wild-type but did not exhibit reduced male fertility. Since 12 genes were not individually essential for male fertilization, it is unlikely that these genes could be the cause of infertility or contraceptive targets. It is better to focus on other essential genes because complementary genes to these 12 genes may exist.
Keywords: CRISPR/Cas9; Knockout mice; Male infertility; Spermatozoa; Testis.
Copyright © 2024 The Authors. Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest All authors declared no competing interests.
Figures
References
-
- Aoki Y, Tsujimura A, Kaseda K, Okabe M, Tokuhiro K, Ohta T, O’Bryan MK, Okuda H, Kitamura K, Ogawa Y, Fujiki T, Wada M, Horie S, Nishimune Y, Tanaka H 2020. Haprin-deficient spermatozoa are incapable of in vitro fertilization. Mol. Reprod. Dev 87 (5), 534–541. - PubMed
-
- Fang Y, Liu D, Yang Y, Zhang H, Wu H, Jiang H, Zhao L, Zhang Z 2020. Altered gene expression profiles of testicular tissues from azoospermic patients with maturation arrest. Andrologia 52 (11), e13812. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

