Tracking antimicrobial resistance transmission in urban and rural communities in Bangladesh: a One Health study of genomic diversity of ESBL-producing and carbapenem-resistant Escherichia coli
- PMID: 38700359
- PMCID: PMC11237648
- DOI: 10.1128/spectrum.03956-23
Tracking antimicrobial resistance transmission in urban and rural communities in Bangladesh: a One Health study of genomic diversity of ESBL-producing and carbapenem-resistant Escherichia coli
Abstract
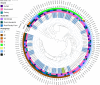
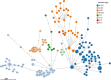
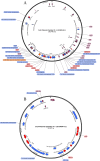
Antimicrobial resistance (AMR) poses a significant threat to global health and sustainable development goals, especially in low- and middle-income countries (LMICs). This study aimed to understand the transmission of AMR between poultry, humans, and the environment in Bangladesh using a One Health approach. We analyzed the whole genome sequences (WGS) of 117 extended-spectrum β-lactamase-producing Escherichia coli (ESBL-Ec) isolates, with 46 being carbapenem resistant. These isolates were obtained from human (n = 20) and poultry feces (n = 12), as well as proximal environments (wastewater) (n = 85) of three different study sites, including rural households (n = 48), rural poultry farms (n = 20), and urban wet markets (n = 49). The WGS of ESBL-Ec isolates were compared with 58 clinical isolates from global databases. No significant differences in antibiotic resistance genes (ARGs) were observed in ESBL-Ec isolated from humans with and without exposure to poultry. Environmental isolates showed higher ARG diversity than human and poultry isolates. No clonal transmission between poultry and human isolates was found, but wastewater was a reservoir for ESBL-Ec for both. Except for one human isolate, all ESBL-Ec isolates were distinct from clinical isolates. Most isolates (77.8%) carried at least one plasmid replicon type, with IncFII being the most prevalent. IncFIA was predominant in human isolates, while IncFII, Col(MG828), and p0111 were common in poultry. We observed putative sharing of ARG-carrying plasmids among isolates, mainly from wastewater. However, in most cases, bacterial isolates sharing plasmids were also clonally related, suggesting clonal spread was more probable than just plasmid transfer.
Importance: Our study underscores that wastewater discharged from households and wet markets carries antibiotic-resistant organisms from both human and animal sources. Thus, direct disposal of wastewater into the environment not only threatens human health but also endangers food safety by facilitating the spread of antimicrobial resistance (AMR) to surface water, crops, vegetables, and subsequently to food-producing animals. In regions with intensive poultry production heavily reliant on the prophylactic use of antibiotics, compounded by inadequate waste management systems, such as Bangladesh, the ramifications are particularly pronounced. Wastewater serves as a pivotal juncture for the dissemination of antibiotic-resistant organisms and functions as a pathway through which strains of human and animal origin can infiltrate the environment and potentially colonize new hosts. Further research is needed to thoroughly characterize wastewater isolates/populations and understand their potential impact on interconnected environments, communities, and wildlife.
Keywords: ESBL-producing E. coli; antibiotic resistance genes; clonal spread; wastewater; whole genome sequencing.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Genomic insights into extended-spectrum β-lactamase- and plasmid-borne AmpC-producing Escherichia coli transmission between humans and livestock in rural Cambodia.J Med Microbiol. 2025 Mar;74(3):001988. doi: 10.1099/jmm.0.001988. J Med Microbiol. 2025. PMID: 40079731 Free PMC article.
-
Genomic characterization of extended-spectrum β-lactamase-producing Escherichia coli spread among chickens and healthy residents in Lombok, Indonesia.Appl Environ Microbiol. 2025 May 21;91(5):e0236424. doi: 10.1128/aem.02364-24. Epub 2025 Apr 11. Appl Environ Microbiol. 2025. PMID: 40214227 Free PMC article.
-
Occurrence of the Colistin Resistance Gene mcr-1 and Additional Antibiotic Resistance Genes in ESBL/AmpC-Producing Escherichia coli from Poultry in Lebanon: A Nationwide Survey.Microbiol Spectr. 2021 Oct 31;9(2):e0002521. doi: 10.1128/Spectrum.00025-21. Epub 2021 Sep 8. Microbiol Spectr. 2021. PMID: 34494875 Free PMC article.
-
Emerging threats: Antimicrobial resistance in extended-spectrum beta-lactamase and carbapenem-resistant Escherichia coli.Microb Pathog. 2025 Mar;200:107275. doi: 10.1016/j.micpath.2024.107275. Epub 2025 Jan 9. Microb Pathog. 2025. PMID: 39798725 Review.
-
Exploring Extended-Spectrum Beta-Lactamase (ESBL)-Producing Escherichia coli in Food-Producing Animals and Animal-Derived Foods.Pathogens. 2024 Apr 22;13(4):346. doi: 10.3390/pathogens13040346. Pathogens. 2024. PMID: 38668301 Free PMC article. Review.
Cited by
-
Limited clonality but widespread plasmid sharing of ESBL-producing E. coli between humans and the environment of northeastern Slovenia.Curr Res Microb Sci. 2025 May 21;9:100408. doi: 10.1016/j.crmicr.2025.100408. eCollection 2025. Curr Res Microb Sci. 2025. PMID: 40511295 Free PMC article.
-
Xenobiotics and Broiler Microbiota: Molecular Insights into Bacterial Antimicrobial Resistance and Food Safety Implications for Human Health.J Xenobiot. 2025 Aug 8;15(4):129. doi: 10.3390/jox15040129. J Xenobiot. 2025. PMID: 40863336 Free PMC article. Review.
References
-
- Devendra C, Chantalakhana C. 2002. Animals, poor people and food insecurity: opportunities for improved livelihoods through efficient natural resource management. Outlook Agric 31:161–175. doi:10.5367/000000002101294010 - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

