Fusion/fission protein family identification in Archaea
- PMID: 38700364
- PMCID: PMC11237513
- DOI: 10.1128/msystems.00948-23
Fusion/fission protein family identification in Archaea
Abstract
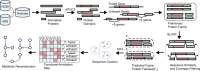
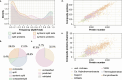
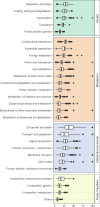
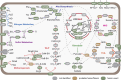
The majority of newly discovered archaeal lineages remain without a cultivated representative, but scarce experimental data from the cultivated organisms show that they harbor distinct functional repertoires. To unveil the ecological as well as evolutionary impact of Archaea from metagenomics, new computational methods need to be developed, followed by in-depth analysis. Among them is the genome-wide protein fusion screening performed here. Natural fusions and fissions of genes not only contribute to microbial evolution but also complicate the correct identification and functional annotation of sequences. The products of these processes can be defined as fusion (or composite) proteins, the ones consisting of two or more domains originally encoded by different genes and split proteins, and the ones originating from the separation of a gene in two (fission). Fusion identifications are required for proper phylogenetic reconstructions and metabolic pathway completeness assessments, while mappings between fused and unfused proteins can fill some of the existing gaps in metabolic models. In the archaeal genome-wide screening, more than 1,900 fusion/fission protein clusters were identified, belonging to both newly sequenced and well-studied lineages. These protein families are mainly associated with different types of metabolism, genetic, and cellular processes. Moreover, 162 of the identified fusion/fission protein families are archaeal specific, having no identified fused homolog within the bacterial domain. Our approach was validated by the identification of experimentally characterized fusion/fission cases. However, around 25% of the identified fusion/fission families lack functional annotations for both composite and split states, showing the need for experimental characterization in Archaea.IMPORTANCEGenome-wide fusion screening has never been performed in Archaea on a broad taxonomic scale. The overlay of multiple computational techniques allows the detection of a fine-grained set of predicted fusion/fission families, instead of rough estimations based on conserved domain annotations only. The exhaustive mapping of fused proteins to bacterial organisms allows us to capture fusion/fission families that are specific to archaeal biology, as well as to identify links between bacterial and archaeal lineages based on cooccurrence of taxonomically restricted proteins and their sequence features. Furthermore, the identification of poorly characterized lineage-specific fusion proteins opens up possibilities for future experimental and computational investigations. This approach enhances our understanding of Archaea in general and provides potential candidates for in-depth studies in the future.
Keywords: archaeal biology; archaeal evolution; bacterial fusions; comparative genomics; large-scale screening.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Genome-wide identification of SF1 and SF2 helicases from archaea.Gene. 2016 Jan 15;576(1 Pt 2):214-28. doi: 10.1016/j.gene.2015.10.007. Epub 2015 Oct 8. Gene. 2016. PMID: 26456193
-
Phyletic Distribution and Lineage-Specific Domain Architectures of Archaeal Two-Component Signal Transduction Systems.J Bacteriol. 2018 Mar 12;200(7):e00681-17. doi: 10.1128/JB.00681-17. Print 2018 Apr 1. J Bacteriol. 2018. PMID: 29263101 Free PMC article.
-
Identification and characterization of archaeal pseudomurein biosynthesis genes through pangenomics.mSystems. 2025 Mar 18;10(3):e0140124. doi: 10.1128/msystems.01401-24. Epub 2025 Feb 12. mSystems. 2025. PMID: 39936904 Free PMC article.
-
Stress genes and proteins in the archaea.Microbiol Mol Biol Rev. 1999 Dec;63(4):923-67, table of contents. doi: 10.1128/MMBR.63.4.923-967.1999. Microbiol Mol Biol Rev. 1999. PMID: 10585970 Free PMC article. Review.
-
Towards functional characterization of archaeal genomic dark matter.Biochem Soc Trans. 2019 Feb 28;47(1):389-398. doi: 10.1042/BST20180560. Epub 2019 Feb 1. Biochem Soc Trans. 2019. PMID: 30710061 Free PMC article. Review.
Cited by
-
Navigating the archaeal frontier: insights and projections from bioinformatic pipelines.Front Microbiol. 2024 Sep 23;15:1433224. doi: 10.3389/fmicb.2024.1433224. eCollection 2024. Front Microbiol. 2024. PMID: 39380680 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

