Kernel-based testing for single-cell differential analysis
- PMID: 38702740
- PMCID: PMC11069218
- DOI: 10.1186/s13059-024-03255-1
Kernel-based testing for single-cell differential analysis
Abstract
Single-cell technologies offer insights into molecular feature distributions, but comparing them poses challenges. We propose a kernel-testing framework for non-linear cell-wise distribution comparison, analyzing gene expression and epigenomic modifications. Our method allows feature-wise and global transcriptome/epigenome comparisons, revealing cell population heterogeneities. Using a classifier based on embedding variability, we identify transitions in cell states, overcoming limitations of traditional single-cell analysis. Applied to single-cell ChIP-Seq data, our approach identifies untreated breast cancer cells with an epigenomic profile resembling persister cells. This demonstrates the effectiveness of kernel testing in uncovering subtle population variations that might be missed by other methods.
Keywords: Differential analysis; Kernel methods; Single cell epigenomics; Single cell transcriptomics.
© 2024. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



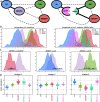

References
-
- Angelidis I, Simon LM, Fernandez IE, Strunz M, Mayr CH, Greiffo FR, Tsitsiridis G, Ansari M, Graf E, Strom T-M, Nagendran M, Desai T, Eickelberg O, Mann M, Theis FJ, Schiller HB. An atlas of the aging lung mapped by single cell transcriptomics and deep tissue proteomics. Nat Commun. 2019;10(1):963. Number: 1 Publisher: Nature Publishing Group. - PMC - PubMed
-
- Bach FR, Lanckriet GRG, Jordan MI. Multiple kernel learning, conic duality, and the SMO algorithm. In: Proceedings of the twenty-first international conference on machine learning, ICML ’04. New York: Association for Computing Machinery; 2004. p. 6
-
- Banerjee T, Bhattacharya BB, Mukherjee G. A nearest-neighbor based nonparametric test for viral remodeling in heterogeneous single-cell proteomic data. Ann Appl Stat. 2020;14(4):1777–1805. doi: 10.1214/20-AOAS1362. - DOI
-
- Benjamini et Hochberg. Controlling the false discovery rate: a practical and powerful approach to multiple testing on JSTOR. 1995.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

