Three steps towards comparability and standardization among molecular methods for characterizing insect communities
- PMID: 38705189
- PMCID: PMC11070264
- DOI: 10.1098/rstb.2023.0118
Three steps towards comparability and standardization among molecular methods for characterizing insect communities
Abstract
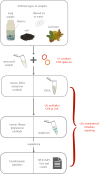
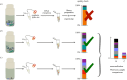
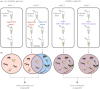
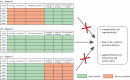
Molecular methods are currently some of the best-suited technologies for implementation in insect monitoring. However, the field is developing rapidly and lacks agreement on methodology or community standards. To apply DNA-based methods in large-scale monitoring, and to gain insight across commensurate data, we need easy-to-implement standards that improve data comparability. Here, we provide three recommendations for how to improve and harmonize efforts in biodiversity assessment and monitoring via metabarcoding: (i) we should adopt the use of synthetic spike-ins, which will act as positive controls and internal standards; (ii) we should consider using several markers through a multiplex polymerase chain reaction (PCR) approach; and (iii) we should commit to the publication and transparency of all protocol-associated metadata in a standardized fashion. For (i), we provide a ready-to-use recipe for synthetic cytochrome c oxidase spike-ins, which enable between-sample comparisons. For (ii), we propose two gene regions for the implementation of multiplex PCR approaches, thereby achieving a more comprehensive community description. For (iii), we offer guidelines for transparent and unified reporting of field, wet-laboratory and dry-laboratory procedures, as a key to making comparisons between studies. Together, we feel that these three advances will result in joint quality and calibration standards rather than the current laboratory-specific proof of concepts. This article is part of the theme issue 'Towards a toolkit for global insect biodiversity monitoring'.
Keywords: COI synthetic spike-in; comparability; internal standard; metabarcoding; metadata; multiplex.
Conflict of interest statement
We declare we have no competing interests.
Figures
Similar articles
-
A shortcut to sample coverage standardization in metabarcoding data provides new insights into land-use effects on insect diversity.Proc Biol Sci. 2025 May;292(2046):20242927. doi: 10.1098/rspb.2024.2927. Epub 2025 May 7. Proc Biol Sci. 2025. PMID: 40328305 Free PMC article.
-
Sharing insect data through GBIF: novel monitoring methods, opportunities and standards.Philos Trans R Soc Lond B Biol Sci. 2024 Jun 24;379(1904):20230104. doi: 10.1098/rstb.2023.0104. Epub 2024 May 6. Philos Trans R Soc Lond B Biol Sci. 2024. PMID: 38705176 Free PMC article.
-
Towards holistic insect monitoring: species discovery, description, identification and traits for all insects.Philos Trans R Soc Lond B Biol Sci. 2024 Jun 24;379(1904):20230120. doi: 10.1098/rstb.2023.0120. Epub 2024 May 6. Philos Trans R Soc Lond B Biol Sci. 2024. PMID: 38705187 Free PMC article. Review.
-
FAVIS: Fast and versatile protocol for non-destructive metabarcoding of bulk insect samples.PLoS One. 2023 Jul 19;18(7):e0286272. doi: 10.1371/journal.pone.0286272. eCollection 2023. PLoS One. 2023. PMID: 37467453 Free PMC article.
-
Future of DNA-based insect monitoring.Trends Genet. 2023 Jul;39(7):531-544. doi: 10.1016/j.tig.2023.02.012. Epub 2023 Mar 10. Trends Genet. 2023. PMID: 36907721 Review.
Cited by
-
Delivering on a promise: futureproofing automated insect monitoring methods.Philos Trans R Soc Lond B Biol Sci. 2024 Jun 24;379(1904):20230105. doi: 10.1098/rstb.2023.0105. Epub 2024 May 6. Philos Trans R Soc Lond B Biol Sci. 2024. PMID: 38705192 Free PMC article.
-
What evidence exists on wild bee trends in Germany? A systematic map.Environ Evid. 2025 Jun 19;14(1):11. doi: 10.1186/s13750-025-00364-7. Environ Evid. 2025. PMID: 40537832 Free PMC article.
-
Measuring the state of aquatic environments using eDNA-upscaling spatial resolution of biotic indices.Philos Trans R Soc Lond B Biol Sci. 2024 Jun 24;379(1904):20230121. doi: 10.1098/rstb.2023.0121. Epub 2024 May 6. Philos Trans R Soc Lond B Biol Sci. 2024. PMID: 38705183 Free PMC article.
-
Towards a toolkit for global insect biodiversity monitoring.Philos Trans R Soc Lond B Biol Sci. 2024 Jun 24;379(1904):20230101. doi: 10.1098/rstb.2023.0101. Epub 2024 May 6. Philos Trans R Soc Lond B Biol Sci. 2024. PMID: 38705179 Free PMC article.
References
-
- Grimaldi D, Engel M. 2005. Evolution of the insects. Cambridge, UK: Cambridge University Press.
-
- Eggleton P. 2020. The state of the World's insects. Annu. Rev. Environ. Resour. 45, 61-82. (10.1146/annurev-environ-012420-050035) - DOI
-
- Jankielsohn A. 2018. The importance of insects in agricultural ecosystems. Adv. Entomol. 6, 62-73. (10.4236/ae.2018.62006) - DOI
-
- Scudder GGE. 2017. The importance of insects. In Insect biodiversity (eds Foottit RG, Adler PH), pp. 9-43. Chichester, UK: John Wiley & Sons, Ltd.
-
- Kim KC. 2017. Taxonomy and management of insect biodiversity. In Insect biodiversity (eds Foottit RG, Adler PH), pp. 767-782. Chichester, UK: John Wiley & Sons, Ltd.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

