Perinuclear damage from nuclear envelope deterioration elicits stress responses that contribute to LMNA cardiomyopathy
- PMID: 38718107
- PMCID: PMC11078192
- DOI: 10.1126/sciadv.adh0798
Perinuclear damage from nuclear envelope deterioration elicits stress responses that contribute to LMNA cardiomyopathy
Abstract
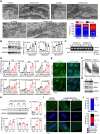
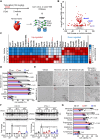
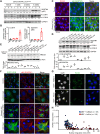
Mutations in the LMNA gene encoding lamins A/C cause an array of tissue-selective diseases, with the heart being the most commonly affected organ. Despite progress in understanding the perturbations emanating from LMNA mutations, an integrative understanding of the pathogenesis underlying cardiac dysfunction remains elusive. Using a novel conditional deletion model capable of translatome profiling, we observed that cardiomyocyte-specific Lmna deletion in adult mice led to rapid cardiomyopathy with pathological remodeling. Before cardiac dysfunction, Lmna-deleted cardiomyocytes displayed nuclear abnormalities, Golgi dilation/fragmentation, and CREB3-mediated stress activation. Translatome profiling identified MED25 activation, a transcriptional cofactor that regulates Golgi stress. Autophagy is disrupted in the hearts of these mice, which can be recapitulated by disrupting the Golgi. Systemic administration of modulators of autophagy or ER stress significantly delayed cardiac dysfunction and prolonged survival. These studies support a hypothesis wherein stress responses emanating from the perinuclear space contribute to the LMNA cardiomyopathy development.
Figures




Update of
-
Perinuclear damage from nuclear envelope deterioration elicits stress responses that contribute to LMNA cardiomyopathy.bioRxiv [Preprint]. 2023 Jun 7:2023.02.14.528563. doi: 10.1101/2023.02.14.528563. bioRxiv. 2023. Update in: Sci Adv. 2024 May 10;10(19):eadh0798. doi: 10.1126/sciadv.adh0798. PMID: 36824975 Free PMC article. Updated. Preprint.
References
-
- Dauer W. T., Worman H. J., The nuclear envelope as a signaling node in development and disease. Dev. Cell 17, 626–638 (2009). - PubMed
-
- Aebi U., Cohn J., Buhle L., Gerace L., The nuclear lamina is a meshwork of intermediate-type filaments. Nature 323, 560–564 (1986). - PubMed
-
- Broers J. L., Peeters E. A., Kuijpers H. J., Endert J., Bouten C. V., Oomens C. W., Baaijens F. P., Ramaekers F. C., Decreased mechanical stiffness in LMNA−/− cells is caused by defective nucleo-cytoskeletal integrity: Implications for the development of laminopathies. Hum. Mol. Genet. 13, 2567–2580 (2004). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

