Rhizobia-diatom symbiosis fixes missing nitrogen in the ocean
- PMID: 38723661
- PMCID: PMC11208148
- DOI: 10.1038/s41586-024-07495-w
Rhizobia-diatom symbiosis fixes missing nitrogen in the ocean
Abstract
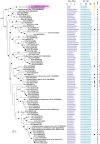
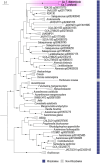
Nitrogen (N2) fixation in oligotrophic surface waters is the main source of new nitrogen to the ocean1 and has a key role in fuelling the biological carbon pump2. Oceanic N2 fixation has been attributed almost exclusively to cyanobacteria, even though genes encoding nitrogenase, the enzyme that fixes N2 into ammonia, are widespread among marine bacteria and archaea3-5. Little is known about these non-cyanobacterial N2 fixers, and direct proof that they can fix nitrogen in the ocean has so far been lacking. Here we report the discovery of a non-cyanobacterial N2-fixing symbiont, 'Candidatus Tectiglobus diatomicola', which provides its diatom host with fixed nitrogen in return for photosynthetic carbon. The N2-fixing symbiont belongs to the order Rhizobiales and its association with a unicellular diatom expands the known hosts for this order beyond the well-known N2-fixing rhizobia-legume symbioses on land6. Our results show that the rhizobia-diatom symbioses can contribute as much fixed nitrogen as can cyanobacterial N2 fixers in the tropical North Atlantic, and that they might be responsible for N2 fixation in the vast regions of the ocean in which cyanobacteria are too rare to account for the measured rates.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






References
-
- Gruber N, Galloway JN. An Earth-system perspective of the global nitrogen cycle. Nature. 2008;451:293–296. - PubMed
-
- Wang W-L, Moore JK, Martiny AC, Primeau FW. Convergent estimates of marine nitrogen fixation. Nature. 2019;566:205–211. - PubMed
-
- Zehr JP, Capone DG. Changing perspectives in marine nitrogen fixation. Science. 2020;368:eaay9514. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

