Viruses contribute to microbial diversification in the rumen ecosystem and are associated with certain animal production traits
- PMID: 38725064
- PMCID: PMC11080232
- DOI: 10.1186/s40168-024-01791-3
Viruses contribute to microbial diversification in the rumen ecosystem and are associated with certain animal production traits
Abstract
Background: The rumen microbiome enables ruminants to digest otherwise indigestible feedstuffs, thereby facilitating the production of high-quality protein, albeit with suboptimal efficiency and producing methane. Despite extensive research delineating associations between the rumen microbiome and ruminant production traits, the functional roles of the pervasive and diverse rumen virome remain to be determined.
Results: Leveraging a recent comprehensive rumen virome database, this study analyzes virus-microbe linkages, at both species and strain levels, across 551 rumen metagenomes, elucidating patterns of microbial and viral diversity, co-occurrence, and virus-microbe interactions. Additionally, this study assesses the potential role of rumen viruses in microbial diversification by analyzing prophages found in rumen metagenome-assembled genomes. Employing CRISPR-Cas spacer-based matching and virus-microbe co-occurrence network analysis, this study suggests that the viruses in the rumen may regulate microbes at strain and community levels through both antagonistic and mutualistic interactions. Moreover, this study establishes that the rumen virome demonstrates responsiveness to dietary shifts and associations with key animal production traits, including feed efficiency, lactation performance, weight gain, and methane emissions.
Conclusions: These findings provide a substantive framework for further investigations to unravel the functional roles of the virome in the rumen in shaping the microbiome and influencing overall animal production performance. Video Abstract.
Keywords: Microbiome; Network analysis; Rumen; Strain diversity; Virome.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


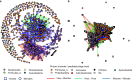



References
-
- Ritchie A, Robinson I, Allison M. Rumen bacteriophage: survey of morphological types. Microscopie Electronique. 1970;3:333–4.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

