Screening and validation of atherosclerosis PAN-apoptotic immune-related genes based on single-cell sequencing
- PMID: 38736872
- PMCID: PMC11082397
- DOI: 10.3389/fimmu.2024.1297298
Screening and validation of atherosclerosis PAN-apoptotic immune-related genes based on single-cell sequencing
Abstract
Background: Carotid atherosclerosis (CAS) is a complication of atherosclerosis (AS). PAN-optosome is an inflammatory programmed cell death pathway event regulated by the PAN-optosome complex. CAS's PAN-optosome-related genes (PORGs) have yet to be studied. Hence, screening the PAN-optosome-related diagnostic genes for treating CAS was vital.
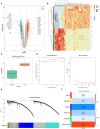
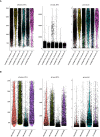
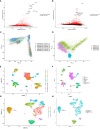
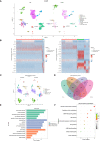
Methods: We introduced transcriptome data to screen out differentially expressed genes (DEGs) in CAS. Subsequently, WGCNA analysis was utilized to mine module genes about PANoptosis score. We performed differential expression analysis (CAS samples vs. standard samples) to obtain CAS-related differentially expressed genes at the single-cell level. Venn diagram was executed to identify PAN-optosome-related differential genes (POR-DEGs) associated with CAS. Further, LASSO regression and RF algorithm were implemented to were executed to build a diagnostic model. We additionally performed immune infiltration and gene set enrichment analysis (GSEA) based on diagnostic genes. We verified the accuracy of the model genes by single-cell nuclear sequencing and RT-qPCR validation of clinical samples, as well as in vitro cellular experiments.
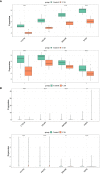
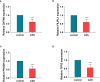
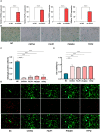
Results: We identified 785 DEGs associated with CAS. Then, 4296 module genes about PANoptosis score were obtained. We obtained the 7365 and 1631 CAS-related DEGs at the single-cell level, respectively. 67 POR-DEGs were retained Venn diagram. Subsequently, 4 PAN-optosome-related diagnostic genes (CNTN4, FILIP1, PHGDH, and TFPI2) were identified via machine learning. Cellular function tests on four genes showed that these genes have essential roles in maintaining arterial cell viability and resisting cellular senescence.
Conclusion: We obtained four PANoptosis-related diagnostic genes (CNTN4, FILIP1, PHGDH, and TFPI2) associated with CAS, laying a theoretical foundation for treating CAS.
Keywords: PAN-apoptosome related genes; bioinformatics; carotid atherosclerosis; diagnostic genes; single-cell sequencing.
Copyright © 2024 Song, Lou, Wang, Zhang, Xia, Ban, Zhao, Sun, Wang, Lin, Guo, Zhou and Xia.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

