Coordinated immune dysregulation in juvenile dermatomyositis revealed by single-cell genomics
- PMID: 38743491
- PMCID: PMC11383589
- DOI: 10.1172/jci.insight.176963
Coordinated immune dysregulation in juvenile dermatomyositis revealed by single-cell genomics
Abstract
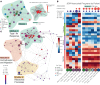
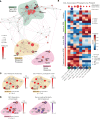
Juvenile dermatomyositis (JDM) is one of several childhood-onset autoimmune disorders characterized by a type I IFN response and autoantibodies. Treatment options are limited due to an incomplete understanding of how the disease emerges from dysregulated cell states across the immune system. We therefore investigated the blood of patients with JDM at different stages of disease activity using single-cell transcriptomics paired with surface protein expression. By immunophenotyping peripheral blood mononuclear cells, we observed skewing of the B cell compartment toward an immature naive state as a hallmark of JDM at diagnosis. Furthermore, we find that these changes in B cells are paralleled by T cell signatures suggestive of Th2-mediated inflammation that persist despite disease quiescence. We applied network analysis to reveal that hyperactivation of the type I IFN response in all immune populations is coordinated with previously masked cell states including dysfunctional protein processing in CD4+ T cells and regulation of cell death programming in NK cells, CD8+ T cells, and γδ T cells. Together, these findings unveil the coordinated immune dysregulation underpinning JDM and provide insight into strategies for restoring balance in immune function.
Keywords: Autoimmune diseases; Autoimmunity; Bioinformatics; Rheumatology.
Figures






Update of
-
Coordinated immune dysregulation in Juvenile Dermatomyositis revealed by single-cell genomics.bioRxiv [Preprint]. 2023 Nov 10:2023.11.07.566033. doi: 10.1101/2023.11.07.566033. bioRxiv. 2023. Update in: JCI Insight. 2024 May 14;9(12):e176963. doi: 10.1172/jci.insight.176963. PMID: 37986917 Free PMC article. Updated. Preprint.
Similar articles
-
Coordinated immune dysregulation in Juvenile Dermatomyositis revealed by single-cell genomics.bioRxiv [Preprint]. 2023 Nov 10:2023.11.07.566033. doi: 10.1101/2023.11.07.566033. bioRxiv. 2023. Update in: JCI Insight. 2024 May 14;9(12):e176963. doi: 10.1172/jci.insight.176963. PMID: 37986917 Free PMC article. Updated. Preprint.
-
Multi-Modal Single-Cell Sequencing Identifies Cellular Immunophenotypes Associated With Juvenile Dermatomyositis Disease Activity.Front Immunol. 2022 Jun 21;13:902232. doi: 10.3389/fimmu.2022.902232. eCollection 2022. Front Immunol. 2022. PMID: 35799782 Free PMC article.
-
Transcriptomes of peripheral blood mononuclear cells from juvenile dermatomyositis patients show elevated inflammation even when clinically inactive.Sci Rep. 2022 Jan 7;12(1):275. doi: 10.1038/s41598-021-04302-8. Sci Rep. 2022. PMID: 34997119 Free PMC article.
-
Immunopathogenesis of juvenile dermatomyositis.Muscle Nerve. 2010 May;41(5):581-92. doi: 10.1002/mus.21669. Muscle Nerve. 2010. PMID: 20405498 Review.
-
Juvenile dermatomyositis: the association of the TNF alpha-308A allele and disease chronicity.Curr Rheumatol Rep. 2001 Oct;3(5):379-86. doi: 10.1007/s11926-996-0007-5. Curr Rheumatol Rep. 2001. PMID: 11564368 Review.
Cited by
-
Decreased Peripheral Blood Natural Killer Cell Count in Untreated Juvenile Dermatomyositis Is Associated with Muscle Weakness.Int J Mol Sci. 2024 Jun 28;25(13):7126. doi: 10.3390/ijms25137126. Int J Mol Sci. 2024. PMID: 39000234 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

