Remodeling of skeletal muscle myosin metabolic states in hibernating mammals
- PMID: 38752835
- PMCID: PMC11098559
- DOI: 10.7554/eLife.94616
Remodeling of skeletal muscle myosin metabolic states in hibernating mammals
Abstract
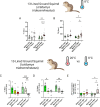
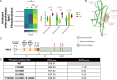
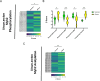
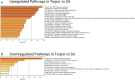
Hibernation is a period of metabolic suppression utilized by many small and large mammal species to survive during winter periods. As the underlying cellular and molecular mechanisms remain incompletely understood, our study aimed to determine whether skeletal muscle myosin and its metabolic efficiency undergo alterations during hibernation to optimize energy utilization. We isolated muscle fibers from small hibernators, Ictidomys tridecemlineatus and Eliomys quercinus and larger hibernators, Ursus arctos and Ursus americanus. We then conducted loaded Mant-ATP chase experiments alongside X-ray diffraction to measure resting myosin dynamics and its ATP demand. In parallel, we performed multiple proteomics analyses. Our results showed a preservation of myosin structure in U. arctos and U. americanus during hibernation, whilst in I. tridecemlineatus and E. quercinus, changes in myosin metabolic states during torpor unexpectedly led to higher levels in energy expenditure of type II, fast-twitch muscle fibers at ambient lab temperatures (20 °C). Upon repeating loaded Mant-ATP chase experiments at 8 °C (near the body temperature of torpid animals), we found that myosin ATP consumption in type II muscle fibers was reduced by 77-107% during torpor compared to active periods. Additionally, we observed Myh2 hyper-phosphorylation during torpor in I. tridecemilineatus, which was predicted to stabilize the myosin molecule. This may act as a potential molecular mechanism mitigating myosin-associated increases in skeletal muscle energy expenditure during periods of torpor in response to cold exposure. Altogether, we demonstrate that resting myosin is altered in hibernating mammals, contributing to significant changes to the ATP consumption of skeletal muscle. Additionally, we observe that it is further altered in response to cold exposure and highlight myosin as a potentially contributor to skeletal muscle non-shivering thermogenesis.
Keywords: biochemistry; cell biology; chemical biology; eliomys quercinus; ictidomys tridecemlineatus; ursus americanus; ursus arctos.
Plain language summary
Many animals use hibernation as a tactic to survive harsh winters. During this dormant, inactive state, animals reduce or limit body processes, such as heart rate and body temperature, to minimise their energy use. To conserve energy during hibernation, animals can use different approaches. For example, garden dormice undergo periodic states of extremely low core temperatures (down to 4–8oC); whereas Eurasian brown bears see milder temperature drops (down to 23–25oC). An important organ that changes during hibernation is skeletal muscle. Skeletal muscle typically uses large amounts of energy, making up around 50% of body mass. To survive, hibernating animals must change how their skeletal muscle uses energy. Traditionally, active myosin – a protein found in muscles that helps muscles to contract – was thought to be responsible for most of the energy use by skeletal muscle. But, more recently, resting myosin has also been found to use energy when muscles are relaxed. Lewis et al. studied myosin and skeletal muscle energy use changes during hibernation and whether they could impact the metabolism of hibernating animals. Lewis et al. assessed myosin changes in muscle samples from squirrels, dormice and bears during hibernation and during activity. Experiments showed changes in resting myosin in squirrels and dormice (whose temperature drops to 4–8oC during hibernation) but not in bears. Further analysis revealed that cooling samples from non-hibernating muscle to 4–8oC increased energy use in resting myosin, thereby generating heat. However, no increase in energy use was found after cooling hibernating muscle samples to 4–8oC. This suggest that resting myosin generates heat at cool temperatures – a mechanism that is switched off in hibernating animals to allow them to cool their body temperature. These findings reveal key insights into how animals conserve energy during hibernation. In addition, the results show that myosin regulates energy use in skeletal muscles, which indicates myosin may be a potential drug target in metabolic diseases, such as obesity.
© 2024, Lewis et al.
Conflict of interest statement
CL, EM, MO, MO, JL, RS, MG, CZ, HI, MK, CM, NA, OF, SG, JS, AG, VF, BB, OT, KD, RS, JO No competing interests declared, AH ALH is an owner of Accelerated Muscle Biotechnologies Consultants LLC, which performed the X-ray data reduction and analysis, but services rendered were not linked to outcome or interpretation
Figures









Update of
-
Remodelling of Skeletal Muscle Myosin Metabolic States in Hibernating Mammals.bioRxiv [Preprint]. 2024 Feb 27:2023.11.14.566992. doi: 10.1101/2023.11.14.566992. bioRxiv. 2024. Update in: Elife. 2024 May 16;13:RP94616. doi: 10.7554/eLife.94616. PMID: 38014200 Free PMC article. Updated. Preprint.
References
-
- Asahi M, Sugita Y, Kurzydlowski K, De Leon S, Tada M, Toyoshima C, MacLennan DH. Sarcolipin regulates sarco(endo)plasmic reticulum Ca2+-ATPase (SERCA) by binding to transmembrane helices alone or in association with phospholamban. PNAS. 2003;100:5040–5045. doi: 10.1073/pnas.0330962100. - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

