Receptor-mediated cargo hitchhiking on bulk autophagy
- PMID: 38755257
- PMCID: PMC11294605
- DOI: 10.1038/s44318-024-00091-8
Receptor-mediated cargo hitchhiking on bulk autophagy
Abstract
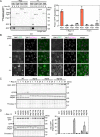
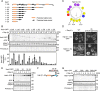
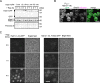
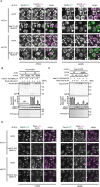
While the molecular mechanism of autophagy is well studied, the cargoes delivered by autophagy remain incompletely characterized. To examine the selectivity of autophagy cargo, we conducted proteomics on isolated yeast autophagic bodies, which are intermediate structures in the autophagy process. We identify a protein, Hab1, that is highly preferentially delivered to vacuoles. The N-terminal 42 amino acid region of Hab1 contains an amphipathic helix and an Atg8-family interacting motif, both of which are necessary and sufficient for the preferential delivery of Hab1 by autophagy. We find that fusion of this region with a cytosolic protein results in preferential delivery of this protein to the vacuole. Furthermore, attachment of this region to an organelle allows for autophagic delivery in a manner independent of canonical autophagy receptor or scaffold proteins. We propose a novel mode of selective autophagy in which a receptor, in this case Hab1, binds directly to forming isolation membranes during bulk autophagy.
Keywords: Saccharomyces cerevisiae; Atg8; Autophagy; Hab1; Selective Autophagy.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures







References
-
- Bindels DS, Haarbosch L, van Weeren L, Postma M, Wiese KE, Mastop M, Aumonier S, Gotthard G, Royant A, Hink MA et al (2017) mScarlet: a bright monomeric red fluorescent protein for cellular imaging. Nat Methods 14:53–56 - PubMed
-
- Drin G, Antonny B (2010) Amphipathic helices and membrane curvature. FEBS Lett 584:1840–7 - PubMed
MeSH terms
Substances
Grants and funding
- 19K16121/MEXT | Japan Society for the Promotion of Science (JSPS)
- 22K15101/MEXT | Japan Society for the Promotion of Science (JSPS)
- 16H06375/MEXT | Japan Society for the Promotion of Science (JSPS)
- 19H05708/MEXT | Japan Society for the Promotion of Science (JSPS)
- 19K06634/MEXT | Japan Society for the Promotion of Science (JSPS)
LinkOut - more resources
Full Text Sources

