Constitutional adaptation to p Ka modulation by remote ester hydrolysis
- PMID: 38756812
- PMCID: PMC11095373
- DOI: 10.1039/d4sc01288g
Constitutional adaptation to p Ka modulation by remote ester hydrolysis
Abstract
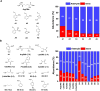
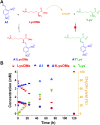
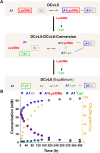
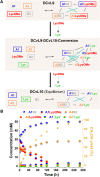
The mechanisms through which environmental conditions affect the expression of interconnected species is a key step to comprehending the principles underlying complex chemical processes. In Nature, chemical modifications triggered by the environment have a major impact on the structure and function of biomolecules and regulate different reaction pathways. Yet, minimalistic artificial systems implementing related adaptation behaviours remain barely explored. The hydrolysis of amino acid methyl esters to their corresponding amino acids leads to a drastic change in pKa (ca. 7 and 9, respectively) that protonates the free amino group at physiological conditions. Dynamic covalent libraries (DCvLs) based on amino acid methyl esters and aldehydes respond to such hydrolysis and lead to constitutional adaptation. Each of the libraries studied experiences a DCvL conversion allowing for constituent selection due to the silencing of the zwitterionic amino acids towards imine formation. The selective action of different enzymes on the DCvLs results in states with simplified constitutional distributions and transient chirality. When additional components (i.e., scavengers) that are not affected by hydrolysis are introduced into the dynamic libraries, the amino acid methyl ester hydrolysis induces the up-regulation of the constituents made of these scavenging components. In these systems, the constituent distribution is resolved from a scrambled mixture of imines to a state characterized by the predominance of a single aldimine. Remarkably, although the final libraries display higher "simplexity", the different transient states present an increased complexity that allows for the emergence of organized structures [micelle formation] and distributions [up-regulation of two antagonistic constituents]. This reactive site inhibition by a remote chemical modification resembles the scenario found in some enzymes for the regulation of their activity through proximal post-translational modifications.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
There are no conflicts to declare.
Figures



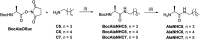

Similar articles
-
Triple Adaptation of Constitutional Dynamic Networks of Imines in Response to Micellar Agents: Internal Uptake-Interfacial Localization-Shape Transition.J Am Chem Soc. 2024 Apr 3;146(13):9096-9111. doi: 10.1021/jacs.3c14200. Epub 2024 Mar 25. J Am Chem Soc. 2024. PMID: 38526415
-
Planning Implications Related to Sterilization-Sensitive Science Investigations Associated with Mars Sample Return (MSR).Astrobiology. 2022 Jun;22(S1):S112-S164. doi: 10.1089/AST.2021.0113. Epub 2022 May 19. Astrobiology. 2022. PMID: 34904892
-
Constitutional Dynamic Selection at Low Reynolds Number in a Triple Dynamic System: Covalent Dynamic Adaptation Driven by Double Supramolecular Self-Assembly.J Am Chem Soc. 2021 Sep 8;143(35):14136-14146. doi: 10.1021/jacs.1c04446. Epub 2021 Aug 25. J Am Chem Soc. 2021. PMID: 34432464
-
Microbial/enzymatic synthesis of chiral drug intermediates.Adv Appl Microbiol. 2000;47:33-78. doi: 10.1016/s0065-2164(00)47001-2. Adv Appl Microbiol. 2000. PMID: 12876794 Review.
-
Constitutional dynamic chemistry: bridge from supramolecular chemistry to adaptive chemistry.Top Curr Chem. 2012;322:1-32. doi: 10.1007/128_2011_256. Top Curr Chem. 2012. PMID: 22169958 Review.
Cited by
-
Chemically Triggered Reactive Coacervates Show Life-Like Budding and Membrane Formation.J Am Chem Soc. 2025 Feb 12;147(6):5293-5299. doi: 10.1021/jacs.4c16416. Epub 2025 Jan 28. J Am Chem Soc. 2025. PMID: 39875119 Free PMC article.
References
-
- Jencks W. P., Catalysis in Chemistry and Enzymology, McGraw-Hill, New York, 1969
LinkOut - more resources
Full Text Sources

