Original COVID-19 priming regimen impacts the immunogenicity of bivalent BA.1 and BA.5 boosters
- PMID: 38762522
- PMCID: PMC11102539
- DOI: 10.1038/s41467-024-48414-x
Original COVID-19 priming regimen impacts the immunogenicity of bivalent BA.1 and BA.5 boosters
Abstract
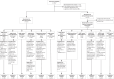
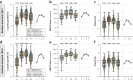
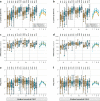
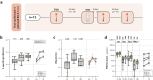
Waning antibody responses after COVID-19 vaccination combined with the emergence of the SARS-CoV-2 Omicron lineage led to reduced vaccine effectiveness. As a countermeasure, bivalent mRNA-based booster vaccines encoding the ancestral spike protein in combination with that of Omicron BA.1 or BA.5 were introduced. Since then, different BA.2-descendent lineages have become dominant, such as XBB.1.5, JN.1, or EG.5.1. Here, we report post-hoc analyses of data from the SWITCH-ON study, assessing how different COVID-19 priming regimens affect the immunogenicity of bivalent booster vaccinations and breakthrough infections (NCT05471440). BA.1 and BA.5 bivalent vaccines boosted neutralizing antibodies and T-cells up to 3 months after boost; however, cross-neutralization of XBB.1.5 was poor. Interestingly, different combinations of prime-boost regimens induced divergent responses: participants primed with Ad26.COV2.S developed lower binding antibody levels after bivalent boost while neutralization and T-cell responses were similar to mRNA-based primed participants. In contrast, the breadth of neutralization was higher in mRNA-primed and bivalent BA.5 boosted participants. Combined, our data further support the current use of monovalent vaccines based on circulating strains when vaccinating risk groups, as recently recommended by the WHO. We emphasize the importance of the continuous assessment of immune responses targeting circulating variants to guide future COVID-19 vaccination policies.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


References
-
- Wu N, et al. Long-term effectiveness of COVID-19 vaccines against infections, hospitalisations, and mortality in adults: findings from a rapid living systematic evidence synthesis and meta-analysis up to December, 2022. Lancet Respir. Med. 2023;11:439–452. doi: 10.1016/S2213-2600(23)00015-2. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
- 10430072110001/ZonMw (Netherlands Organisation for Health Research and Development)
- 10430072110008/ZonMw (Netherlands Organisation for Health Research and Development)
- 10430072110001/ZonMw (Netherlands Organisation for Health Research and Development)
- 10430072110001/ZonMw (Netherlands Organisation for Health Research and Development)
- 10430072110008/ZonMw (Netherlands Organisation for Health Research and Development)
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

