Topographic analysis of pancreatic cancer by TMA and digital spatial profiling reveals biological complexity with potential therapeutic implications
- PMID: 38762572
- PMCID: PMC11102543
- DOI: 10.1038/s41598-024-62031-0
Topographic analysis of pancreatic cancer by TMA and digital spatial profiling reveals biological complexity with potential therapeutic implications
Abstract
Pancreatic ductal adenocarcinoma (PDAC) remains one of the most lethal human malignancies. Tissue microarrays (TMA) are an established method of high throughput biomarker interrogation in tissues but may not capture histological features of cancer with potential biological relevance. Topographic TMAs (T-TMAs) representing pathophysiological hallmarks of cancer were constructed from representative, retrospective PDAC diagnostic material, including 72 individual core tissue samples. The T-TMA was interrogated with tissue hybridization-based experiments to confirm the accuracy of the topographic sampling, expression of pro-tumourigenic and immune mediators of cancer, totalling more than 750 individual biomarker analyses. A custom designed Next Generation Sequencing (NGS) panel and a spatial distribution-specific transcriptomic evaluation were also employed. The morphological choice of the pathophysiological hallmarks of cancer was confirmed by protein-specific expression. Quantitative analysis identified topography-specific patterns of expression in the IDO/TGF-β axis; with a heterogeneous relationship of inflammation and desmoplasia across hallmark areas and a general but variable protein and gene expression of c-MET. NGS results highlighted underlying genetic heterogeneity within samples, which may have a confounding influence on the expression of a particular biomarker. T-TMAs, integrated with quantitative biomarker digital scoring, are useful tools to identify hallmark specific expression of biomarkers in pancreatic cancer.
Keywords: Biomarkers; Digital spatial profiling; Image analysis; Pancreatic ductal adenocarcinoma; Topographic tissue microarrays; Tumour heterogeneity.
© 2024. The Author(s).
Conflict of interest statement
MST is a scientific advisor to Mindpeak and Sonrai Analytics, and has received honoraria recently from BMS, MSD and Incyte. None of these disclosures are related to this work. The remaining authors declare no potential conflicts of interest.
Figures

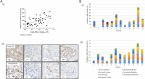

References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

