This is a preprint.
Environment by environment interactions (ExE) differ across genetic backgrounds (ExExG)
- PMID: 38766025
- PMCID: PMC11100745
- DOI: 10.1101/2024.05.08.593194
Environment by environment interactions (ExE) differ across genetic backgrounds (ExExG)
Abstract
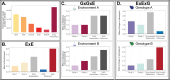
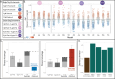
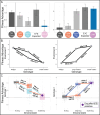
While the terms "gene-by-gene interaction" (GxG) and "gene-by-environment interaction" (GxE) are widely recognized in the fields of quantitative and evolutionary genetics, "environment-byenvironment interaction" (ExE) is a term used less often. In this study, we find that environmentby-environment interactions are a meaningful driver of phenotypes, and moreover, that they differ across different genotypes (suggestive of ExExG). To support this conclusion, we analyzed a large dataset of roughly 1,000 mutant yeast strains with varying degrees of resistance to different antifungal drugs. Our findings reveal that the effectiveness of a drug combination, relative to single drugs, often differs across drug resistant mutants. Remarkably, even mutants that differ by only a single nucleotide change can have dramatically different drug × drug (ExE) interactions. We also introduce a new framework that more accurately predicts the direction and magnitude of ExE interactions for some mutants. Understanding how ExE interactions change across genotypes (ExExG) is crucial not only for modeling the evolution of pathogenic microbes, but also for enhancing our knowledge of the underlying cell biology and the sources of phenotypic variance within populations. While the significance of ExExG interactions has been overlooked in evolutionary and population genetics, these fields and others stand to benefit from understanding how these interactions shape the complex behavior of living systems.
Figures


Similar articles
-
Erratum: High-Throughput Identification of Resistance to Pseudomonas syringae pv. Tomato in Tomato using Seedling Flood Assay.J Vis Exp. 2023 Oct 18;(200). doi: 10.3791/6576. J Vis Exp. 2023. PMID: 37851522
-
Distinguishing mutants that resist drugs via different mechanisms by examining fitness tradeoffs.Elife. 2024 Sep 10;13:RP94144. doi: 10.7554/eLife.94144. Elife. 2024. PMID: 39255191 Free PMC article.
-
Allometric plasticity and the evolution of environment-by-environment (E×E) interactions during a rapid range expansion of a dung beetle.Evolution. 2023 Mar 1;77(3):682-689. doi: 10.1093/evolut/qpac071. Evolution. 2023. PMID: 36626800
-
The genomic determinants of genotype × environment interactions in gene expression.Trends Genet. 2013 Aug;29(8):479-87. doi: 10.1016/j.tig.2013.05.006. Epub 2013 Jun 13. Trends Genet. 2013. PMID: 23769209 Review.
-
Genetic and plastic rewiring of food webs under climate change.J Anim Ecol. 2021 Aug;90(8):1814-1830. doi: 10.1111/1365-2656.13541. Epub 2021 Jun 22. J Anim Ecol. 2021. PMID: 34028791 Free PMC article. Review.
References
-
- Bateson W., Mendel’s principles of heredity (University Press, 1909).
-
- Eguchi Y., Bilolikar G., Geiler-Samerotte K., Why and how to study genetic changes with context-dependent effects. Curr. Opin. Genet. Dev. 58–59, 95–102 (2019). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
