This is a preprint.
Alternative splicing of clock transcript mediates the response of circadian clocks to temperature changes
- PMID: 38766142
- PMCID: PMC11100826
- DOI: 10.1101/2024.05.10.593646
Alternative splicing of clock transcript mediates the response of circadian clocks to temperature changes
Update in
-
Alternative splicing of Clock transcript mediates the response of circadian clocks to temperature changes.Proc Natl Acad Sci U S A. 2024 Dec 10;121(50):e2410680121. doi: 10.1073/pnas.2410680121. Epub 2024 Dec 4. Proc Natl Acad Sci U S A. 2024. PMID: 39630861 Free PMC article.
Abstract
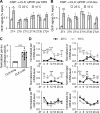
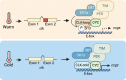
Circadian clocks respond to temperature changes over the calendar year, allowing organisms to adjust their daily biological rhythms to optimize health and fitness. In Drosophila, seasonal adaptations and temperature compensation are regulated by temperature-sensitive alternative splicing (AS) of period (per) and timeless (tim) genes that encode key transcriptional repressors of clock gene expression. Although clock (clk) gene encodes the critical activator of clock gene expression, AS of its transcripts and its potential role in temperature regulation of clock function have not been explored. We therefore sought to investigate whether clk exhibits AS in response to temperature and the functional changes of the differentially spliced transcripts. We observed that clk transcripts indeed undergo temperature-sensitive AS. Specifically, cold temperature leads to the production of an alternative clk transcript, hereinafter termed clk-cold, which encodes a CLK isoform with an in-frame deletion of four amino acids proximal to the DNA binding domain. Notably, serine 13 (S13), which we found to be a CK1α-dependent phosphorylation site, is among the four amino acids deleted in CLK-cold protein. Using a combination of transgenic fly, tissue culture, and in vitro experiments, we demonstrated that upon phosphorylation at CLK(S13), CLK-DNA interaction is reduced, thus decreasing CLK occupancy at clock gene promoters. This is in agreement with our findings that CLK occupancy at clock genes and transcriptional output are elevated at cold temperature, which can be explained by the higher amounts of CLK-cold isoforms that lack S13 residue. This study provides new insights into the complex collaboration between AS and phospho-regulation in shaping temperature responses of the circadian clock.
Keywords: Circadian clock; DNA-binding; alternative splicing; casein kinase 1α; phosphorylation; posttranslational modification; temperature; transcriptional activity.
Conflict of interest statement
Declaration Of Interests The authors declare no competing interests.
Figures





References
-
- Patke A., Young M. W. & Axelrod S. Molecular mechanisms and physiological importance of circadian rhythms. Nat. Rev. Mol. Cell Biol. 21, 67–84 (2020). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
