Construction and Validation of Novel Ferroptosis-related Risk Score Signature and Prognostic Prediction Nomogram for Patients with Colorectal Cancer
- PMID: 38774759
- PMCID: PMC11103399
- DOI: 10.7150/ijms.91446
Construction and Validation of Novel Ferroptosis-related Risk Score Signature and Prognostic Prediction Nomogram for Patients with Colorectal Cancer
Abstract
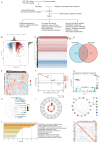
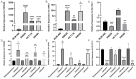
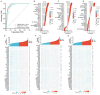
Background: Colorectal cancer (CRC) has a high morbidity and mortality. Ferroptosis is a phenomenon in which metabolism and cell death are closely related. The role of ferroptosis-related genes in the progression of CRC is still not clear. Therefore, we screened and validated the ferroptosis-related genes which could determine the prevalence, risk and prognosis of patients with CRC. Methods: We firstly screened differentially expressed ferroptosis-related genes by The Cancer Genome Atlas (TCGA) database. Then, these genes were used to construct a risk-score model using the least absolute shrinkage and selection operator (LASSO) regression algorithm. The function and prognosis of the ferroptosis-related genes were confirmed using multi-omics analysis. The gene expression results were validated using publicly available databases and qPCR. We also used publicly available data and ferroptosis-related genes to construct a prognostic prediction nomogram. Results: A total of 24 differential expressed genes associated with ferroptosis were screened in this study. A three-gene risk score model was then established based on these 24 genes and GPX3, CDKN2A and SLC7A11 were selected. The significant prognostic value of this novel three-gene signature was also assessed. Furthermore, we conducted RT-qPCR analysis on cell lines and tissues, and validated the high expression of CDKN2A, GPX3 and low expression of SLC7A11 in CRC cells. The observed mRNA expression of GPX3, CDKN2A and SLC7A11 was consistent with the predicted outcomes. Besides, eight variables including selected ferroptosis related genes were included to establish the prognostic prediction nomogram for patients with CRC. The calibration plots showed favorable consistency between the prediction of the nomogram and actual observations. Also, the time-dependent AUC (>0.7) indicated satisfactory discriminative ability of the nomogram. Conclusions: The present study constructed and validated a novel ferroptosis-related three-gene risk score signature and a prognostic prediction nomogram for patients with CRC. Also, we screened and validated the ferroptosis-related genes GPX3, CDKN2A, and SLC7A11 which could serve as novel biomarkers for patients with CRC.
Keywords: Colorectal Cancer; Ferroptosis; Nomogram; Prognosis; Risk model.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures



References
-
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30. - PubMed
-
- Siegel RL, Miller KD, Goding Sauer A, Fedewa SA, Butterly LF, Anderson JC. et al. Colorectal cancer statistics, 2020. CA Cancer J Clin. 2020;70:145–64. - PubMed
-
- Biller LH, Schrag D. Diagnosis and Treatment of Metastatic Colorectal Cancer: A Review. JAMA. 2021;325:669–85. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

