Dating Ammonia-Oxidizing Bacteria with Abundant Eukaryotic Fossils
- PMID: 38776415
- PMCID: PMC11135946
- DOI: 10.1093/molbev/msae096
Dating Ammonia-Oxidizing Bacteria with Abundant Eukaryotic Fossils
Abstract
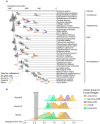
Evolution of a complete nitrogen (N) cycle relies on the onset of ammonia oxidation, which aerobically converts ammonia to nitrogen oxides. However, accurate estimation of the antiquity of ammonia-oxidizing bacteria (AOB) remains challenging because AOB-specific fossils are absent and bacterial fossils amenable to calibrate molecular clocks are rare. Leveraging the ancient endosymbiosis of mitochondria and plastid, as well as using state-of-the-art Bayesian sequential dating approach, we obtained a timeline of AOB evolution calibrated largely by eukaryotic fossils. We show that the first AOB evolved in marine Gammaproteobacteria (Gamma-AOB) and emerged between 2.1 and 1.9 billion years ago (Ga), thus postdating the Great Oxidation Event (GOE; 2.4 to 2.32 Ga). To reconcile the sedimentary N isotopic signatures of ammonia oxidation occurring near the GOE, we propose that ammonia oxidation likely occurred at the common ancestor of Gamma-AOB and Gammaproteobacterial methanotrophs, or the actinobacterial/verrucomicrobial methanotrophs which are known to have ammonia oxidation activities. It is also likely that nitrite was transported from the terrestrial habitats where ammonia oxidation by archaea took place. Further, we show that the Gamma-AOB predated the anaerobic ammonia-oxidizing (anammox) bacteria, implying that the emergence of anammox was constrained by the availability of dedicated ammonia oxidizers which produce nitrite to fuel anammox. Our work supports a new hypothesis that N redox cycle involving nitrogen oxides evolved rather late in the ocean.
Keywords: Great Oxidation Event; ammonia-oxidizing bacteria; molecular clock.
© The Author(s) 2024. Published by Oxford University Press on behalf of Society for Molecular Biology and Evolution.
Conflict of interest statement
Conflict of Interest The authors declare that they have no conflict of interest.
Figures



Similar articles
-
Mimicking the oxygen minimum zones: stimulating interaction of aerobic archaeal and anaerobic bacterial ammonia oxidizers in a laboratory-scale model system.Environ Microbiol. 2012 Dec;14(12):3146-58. doi: 10.1111/j.1462-2920.2012.02894.x. Epub 2012 Oct 12. Environ Microbiol. 2012. PMID: 23057688 Free PMC article.
-
Phanerozoic radiation of ammonia oxidizing bacteria.Sci Rep. 2021 Jan 22;11(1):2070. doi: 10.1038/s41598-021-81718-2. Sci Rep. 2021. PMID: 33483596 Free PMC article.
-
Competition between Ammonia-Oxidizing Archaea and Bacteria from Freshwater Environments.Appl Environ Microbiol. 2021 Sep 28;87(20):e0103821. doi: 10.1128/AEM.01038-21. Epub 2021 Aug 4. Appl Environ Microbiol. 2021. PMID: 34347515 Free PMC article.
-
Insights into the physiology of ammonia-oxidizing microorganisms.Curr Opin Chem Biol. 2019 Apr;49:9-15. doi: 10.1016/j.cbpa.2018.09.003. Epub 2018 Sep 17. Curr Opin Chem Biol. 2019. PMID: 30236860 Review.
-
Research advances of ammonia oxidation microorganisms in wastewater: metabolic characteristics, microbial community, influencing factors and process applications.Bioprocess Biosyst Eng. 2023 May;46(5):621-633. doi: 10.1007/s00449-023-02866-5. Epub 2023 Mar 29. Bioprocess Biosyst Eng. 2023. PMID: 36988685 Review.
Cited by
-
Anomalous δ15N values in the Neoarchean associated with an abundant supply of hydrothermal ammonium.Nat Commun. 2025 Feb 22;16(1):1873. doi: 10.1038/s41467-025-57091-3. Nat Commun. 2025. PMID: 39984454 Free PMC article.
References
-
- Ader M, Thomazo C, Sansjofre P, Busigny V, Papineau D, Laffont R, Cartigny P, Halverson GP. Interpretation of the nitrogen isotopic composition of Precambrian sedimentary rocks: assumptions and perspectives. Chem Geol. 2016:429:93–110. 10.1016/j.chemgeo.2016.02.010. - DOI
MeSH terms
Substances
Grants and funding
- 14107823/Hong Kong Research Grants Council (RGC) General Research Fund
- 42293294/Natural Science Foundation of China
- AoE/M-403/16/Hong Kong Research Grants Council Area of Excellence Scheme
- 2022A1515010844/Guangdong Basic and Applied Basic Research Foundation
- 2021M702296/China Postdoctoral Science Foundation
LinkOut - more resources
Full Text Sources

