Alarmin S100A8 imparts chemoresistance of esophageal cancer by reprogramming cancer-associated fibroblasts
- PMID: 38776909
- PMCID: PMC11228400
- DOI: 10.1016/j.xcrm.2024.101576
Alarmin S100A8 imparts chemoresistance of esophageal cancer by reprogramming cancer-associated fibroblasts
Abstract
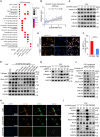
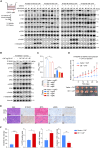
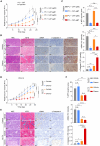
Chemotherapy remains the first-line treatment for advanced esophageal cancer. However, durable benefits are achieved by only a limited subset of individuals due to the elusive chemoresistance. Here, we utilize patient-derived xenografts (PDXs) from esophageal squamous-cell carcinoma to investigate chemoresistance mechanisms in preclinical settings. We observe that activated cancer-associated fibroblasts (CAFs) are enriched in the tumor microenvironment of PDXs resistant to chemotherapy. Mechanistically, we reveal that cancer-cell-derived S100A8 triggers the intracellular RhoA-ROCK-MLC2-MRTF-A pathway by binding to the CD147 receptor of CAFs, inducing CAF polarization and leading to chemoresistance. Therapeutically, we demonstrate that blocking the S100A8-CD147 pathway can improve chemotherapy efficiency. Prognostically, we found the S100A8 levels in peripheral blood can serve as an indicator of chemotherapy responsiveness. Collectively, our study offers a comprehensive understanding of the molecular mechanisms underlying chemoresistance in esophageal cancer and highlights the potential value of S100A8 in the clinical management of esophageal cancer.
Keywords: S100A8; cancer-associated fibroblasts; chemotherapy; esophageal squamous-cell carcinoma; patient-derived xenografts; the tumor microenvironment.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures





References
-
- Matz M., Valkov M., Šekerija M., Luttman S., Caldarella A., Coleman M.P., Allemani C. Worldwide trends in esophageal cancer survival, by sub-site, morphology, and sex: an analysis of 696,974 adults diagnosed in 60 countries during 2000-2014 (CONCORD-3) Cancer Commun. (London, England) 2023;43:963–980. doi: 10.1002/cac2.12457. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

