Rapid DNA unwinding accelerates genome editing by engineered CRISPR-Cas9
- PMID: 38781968
- PMCID: PMC11658890
- DOI: 10.1016/j.cell.2024.04.031
Rapid DNA unwinding accelerates genome editing by engineered CRISPR-Cas9
Abstract
Thermostable clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated (Cas9) enzymes could improve genome-editing efficiency and delivery due to extended protein lifetimes. However, initial experimentation demonstrated Geobacillus stearothermophilus Cas9 (GeoCas9) to be virtually inactive when used in cultured human cells. Laboratory-evolved variants of GeoCas9 overcome this natural limitation by acquiring mutations in the wedge (WED) domain that produce >100-fold-higher genome-editing levels. Cryoelectron microscopy (cryo-EM) structures of the wild-type and improved GeoCas9 (iGeoCas9) enzymes reveal extended contacts between the WED domain of iGeoCas9 and DNA substrates. Biochemical analysis shows that iGeoCas9 accelerates DNA unwinding to capture substrates under the magnesium-restricted conditions typical of mammalian but not bacterial cells. These findings enabled rational engineering of other Cas9 orthologs to enhance genome-editing levels, pointing to a general strategy for editing enzyme improvement. Together, these results uncover a new role for the Cas9 WED domain in DNA unwinding and demonstrate how accelerated target unwinding dramatically improves Cas9-induced genome-editing activity.
Keywords: CRISPR-Cas; Cas9 engineering; DNA unwinding; GeoCas9; R-loop formation; WED domain; cryo-EM; genome editing; iGeoCas9; magnesium.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The Regents of the University of California have patents issued and pending for CRISPR technologies on which the authors are inventors. J.A.D. is a cofounder of Azalea Therapuetics, Caribou Biosciences, Editas Medicine, Evercrisp, Scribe Therapeutics, Intellia Therapeutics, and Mammoth Biosciences. J.A.D. is a scientific advisory board member at Evercrisp, Caribou Biosciences, Intellia Therapeutics, Scribe Therapeutics, Mammoth Biosciences, The Column Group, and Inari. J.A.D. is Chief Science Advisor to Sixth Street; is a Director at Johnson & Johnson, Altos, and Tempus; and has a research project sponsored by Apple Tree Partners.
Figures





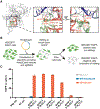
Update of
-
Rapid DNA unwinding accelerates genome editing by engineered CRISPR-Cas9.bioRxiv [Preprint]. 2023 Dec 15:2023.12.14.571777. doi: 10.1101/2023.12.14.571777. bioRxiv. 2023. Update in: Cell. 2024 Jun 20;187(13):3249-3261.e14. doi: 10.1016/j.cell.2024.04.031. PMID: 38168238 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

