Identification of a dCache-type chemoreceptor in Campylobacter jejuni that specifically mediates chemotaxis towards methyl pyruvate
- PMID: 38784811
- PMCID: PMC11111895
- DOI: 10.3389/fmicb.2024.1400284
Identification of a dCache-type chemoreceptor in Campylobacter jejuni that specifically mediates chemotaxis towards methyl pyruvate
Abstract
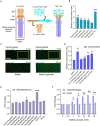
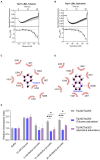
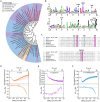
The foodborne pathogenic bacterium Campylobacter jejuni utilizes chemotaxis to assist in the colonization of host niches. A key to revealing the relationship among chemotaxis and pathogenicity is the discovery of signaling molecules perceived by the chemoreceptors. The C. jejuni chemoreceptor Tlp11 is encoded by the highly infective C. jejuni strains. In the present study, we report that the dCache-type ligand-binding domain (LBD) of C. jejuni ATCC 33560 Tlp11 binds directly to novel ligands methyl pyruvate, toluene, and quinoline using the same pocket. Methyl pyruvate elicits a strong chemoattractant response, while toluene and quinoline function as the antagonists without triggering chemotaxis. The sensory LBD was used to control heterologous proteins by constructing chimeras, indicating that the signal induced by methyl pyruvate is transmitted across the membrane. In addition, bioinformatics and experiments revealed that the dCache domains with methyl pyruvate-binding sites and ability are widely distributed in the order Campylobacterales. This is the first report to identify the class of dCache chemoreceptors that bind to attractant methyl pyruvate and antagonists toluene and quinoline. Our research provides a foundation for understanding the chemotaxis and virulence of C. jejuni and lays a basis for the control of this foodborne pathogen.
Keywords: Campylobacter jejuni; chemoreceptor Tlp11; chemotaxis; chimeras; ligands.
Copyright © 2024 Zhao, Yao, Li, Liu and Bi.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
The dCache Domain of the Chemoreceptor Tlp1 in Campylobacter jejuni Binds and Triggers Chemotaxis toward Formate.mBio. 2023 Jun 27;14(3):e0356422. doi: 10.1128/mbio.03564-22. Epub 2023 Apr 13. mBio. 2023. PMID: 37052512 Free PMC article.
-
Diverse Sensory Repertoire of Paralogous Chemoreceptors Tlp2, Tlp3, and Tlp4 in Campylobacter jejuni.Microbiol Spectr. 2022 Dec 21;10(6):e0364622. doi: 10.1128/spectrum.03646-22. Epub 2022 Nov 14. Microbiol Spectr. 2022. PMID: 36374080 Free PMC article.
-
Structure-Activity Relationship Study Reveals the Molecular Basis for Specific Sensing of Hydrophobic Amino Acids by the Campylobacter jejuni Chemoreceptor Tlp3.Biomolecules. 2020 May 11;10(5):744. doi: 10.3390/biom10050744. Biomolecules. 2020. PMID: 32403336 Free PMC article.
-
Campylobacter jejuni transducer like proteins: Chemotaxis and beyond.Gut Microbes. 2017 Jul 4;8(4):323-334. doi: 10.1080/19490976.2017.1279380. Epub 2017 Jan 12. Gut Microbes. 2017. PMID: 28080213 Free PMC article. Review.
-
Chemotaxis in Campylobacter jejuni.Eur J Microbiol Immunol (Bp). 2012 Mar;2(1):24-31. doi: 10.1556/EuJMI.2.2012.1.5. Epub 2012 Mar 17. Eur J Microbiol Immunol (Bp). 2012. PMID: 24611118 Free PMC article. Review.
Cited by
-
Thermal shift assay to identify ligands for bacterial sensor proteins.FEMS Microbiol Rev. 2025 Jan 14;49:fuaf033. doi: 10.1093/femsre/fuaf033. FEMS Microbiol Rev. 2025. PMID: 40720780 Free PMC article. Review.
-
Structural and functional diversity of sensor domains in bacterial transmembrane receptors.Trends Microbiol. 2025 Jul;33(7):796-809. doi: 10.1016/j.tim.2025.02.019. Epub 2025 Mar 21. Trends Microbiol. 2025. PMID: 40121131 Review.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

