When Two Eyes Don't Suffice-Learning Difficult Hyperfluorescence Segmentations in Retinal Fundus Autofluorescence Images via Ensemble Learning
- PMID: 38786570
- PMCID: PMC11122615
- DOI: 10.3390/jimaging10050116
When Two Eyes Don't Suffice-Learning Difficult Hyperfluorescence Segmentations in Retinal Fundus Autofluorescence Images via Ensemble Learning
Abstract
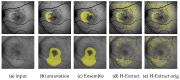
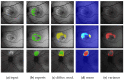
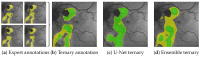
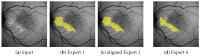
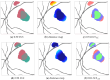
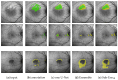
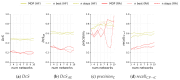
Hyperfluorescence (HF) and reduced autofluorescence (RA) are important biomarkers in fundus autofluorescence images (FAF) for the assessment of health of the retinal pigment epithelium (RPE), an important indicator of disease progression in geographic atrophy (GA) or central serous chorioretinopathy (CSCR). Autofluorescence images have been annotated by human raters, but distinguishing biomarkers (whether signals are increased or decreased) from the normal background proves challenging, with borders being particularly open to interpretation. Consequently, significant variations emerge among different graders, and even within the same grader during repeated annotations. Tests on in-house FAF data show that even highly skilled medical experts, despite previously discussing and settling on precise annotation guidelines, reach a pair-wise agreement measured in a Dice score of no more than 63-80% for HF segmentations and only 14-52% for RA. The data further show that the agreement of our primary annotation expert with herself is a 72% Dice score for HF and 51% for RA. Given these numbers, the task of automated HF and RA segmentation cannot simply be refined to the improvement in a segmentation score. Instead, we propose the use of a segmentation ensemble. Learning from images with a single annotation, the ensemble reaches expert-like performance with an agreement of a 64-81% Dice score for HF and 21-41% for RA with all our experts. In addition, utilizing the mean predictions of the ensemble networks and their variance, we devise ternary segmentations where FAF image areas are labeled either as confident background, confident HF, or potential HF, ensuring that predictions are reliable where they are confident (97% Precision), while detecting all instances of HF (99% Recall) annotated by all experts.
Keywords: CSCR; U-Net; ambiguous; annotation; central serous chorioretinopathy; deep learning; ensemble; fundus autofluorescence; hyperfluorescence; image analysis; inter-observer variability; intra-observer variability; reduced autofluorescence; retinal; segmentation; ternary.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures









References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

