Machine Learning Approach to Metabolomic Data Predicts Type 2 Diabetes Mellitus Incidence
- PMID: 38791370
- PMCID: PMC11120685
- DOI: 10.3390/ijms25105331
Machine Learning Approach to Metabolomic Data Predicts Type 2 Diabetes Mellitus Incidence
Abstract
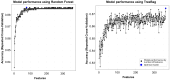
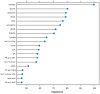
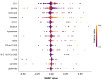
Metabolomics, with its wealth of data, offers a valuable avenue for enhancing predictions and decision-making in diabetes. This observational study aimed to leverage machine learning (ML) algorithms to predict the 4-year risk of developing type 2 diabetes mellitus (T2DM) using targeted quantitative metabolomics data. A cohort of 279 cardiovascular risk patients who underwent coronary angiography and who were initially free of T2DM according to American Diabetes Association (ADA) criteria was analyzed at baseline, including anthropometric data and targeted metabolomics, using liquid chromatography (LC)-mass spectroscopy (MS) and flow injection analysis (FIA)-MS, respectively. All patients were followed for four years. During this time, 11.5% of the patients developed T2DM. After data preprocessing, 362 variables were used for ML, employing the Caret package in R. The dataset was divided into training and test sets (75:25 ratio) and we used an oversampling approach to address the classifier imbalance of T2DM incidence. After an additional recursive feature elimination step, identifying a set of 77 variables that were the most valuable for model generation, a Support Vector Machine (SVM) model with a linear kernel demonstrated the most promising predictive capabilities, exhibiting an F1 score of 50%, a specificity of 93%, and balanced and unbalanced accuracies of 72% and 88%, respectively. The top-ranked features were bile acids, ceramides, amino acids, and hexoses, whereas anthropometric features such as age, sex, waist circumference, or body mass index had no contribution. In conclusion, ML analysis of metabolomics data is a promising tool for identifying individuals at risk of developing T2DM and opens avenues for personalized and early intervention strategies.
Keywords: ML; accuracy; artificial intelligence; diabetes; incidence; machine learning; metabolomics; support vector machine.
Conflict of interest statement
No potential conflicts of interest relevant to this article are reported by A.L., A.M. (Axel Muendlein), S.M., C.H.S., A.M. (Arthur Mader), A.F., P.F., and H.D.
Figures
Similar articles
-
Enhancing type 2 diabetes mellitus prediction by integrating metabolomics and tree-based boosting approaches.Front Endocrinol (Lausanne). 2024 Nov 11;15:1444282. doi: 10.3389/fendo.2024.1444282. eCollection 2024. Front Endocrinol (Lausanne). 2024. PMID: 39588339 Free PMC article.
-
Predictive model and risk analysis for peripheral vascular disease in type 2 diabetes mellitus patients using machine learning and shapley additive explanation.Front Endocrinol (Lausanne). 2024 Feb 28;15:1320335. doi: 10.3389/fendo.2024.1320335. eCollection 2024. Front Endocrinol (Lausanne). 2024. PMID: 38481447 Free PMC article.
-
Predicting long-term type 2 diabetes with support vector machine using oral glucose tolerance test.PLoS One. 2019 Dec 11;14(12):e0219636. doi: 10.1371/journal.pone.0219636. eCollection 2019. PLoS One. 2019. PMID: 31826018 Free PMC article.
-
Metabolomics-Based Prospective Studies and Prediction of Type 2 Diabetes Mellitus Risks.Metab Syndr Relat Disord. 2020 Feb;18(1):1-9. doi: 10.1089/met.2019.0047. Epub 2019 Oct 21. Metab Syndr Relat Disord. 2020. PMID: 31634052 Review.
-
Evaluating Machine Learning Methods of Analyzing Multiclass Metabolomics.J Chem Inf Model. 2023 Dec 25;63(24):7628-7641. doi: 10.1021/acs.jcim.3c01525. Epub 2023 Dec 11. J Chem Inf Model. 2023. PMID: 38079572 Review.
Cited by
-
Special Issue "Machine Learning and Bioinformatics in Human Health and Disease"-Chances and Challenges.Int J Mol Sci. 2024 Nov 28;25(23):12811. doi: 10.3390/ijms252312811. Int J Mol Sci. 2024. PMID: 39684521 Free PMC article.
-
Identification of novel diagnostic biomarkers associated with liver metastasis in colon adenocarcinoma by machine learning.Discov Oncol. 2024 Oct 10;15(1):542. doi: 10.1007/s12672-024-01398-y. Discov Oncol. 2024. PMID: 39390264 Free PMC article.
-
Bioinformatics identification of key microRNA-correlated genes associated with hepatocellular carcinoma heterogeneity and prognosis.BMC Gastroenterol. 2025 Jul 1;25(1):452. doi: 10.1186/s12876-025-04031-6. BMC Gastroenterol. 2025. PMID: 40596892 Free PMC article.
-
Ceramides in cardiovascular disease: emerging role as independent risk predictors and novel therapeutic targets.Cardiovasc Res. 2025 Aug 14;121(9):1345-1358. doi: 10.1093/cvr/cvaf093. Cardiovasc Res. 2025. PMID: 40460239 Free PMC article. Review.
-
Metabolomics: Uncovering Insights into Obesity and Diabetes.Int J Mol Sci. 2025 Jun 27;26(13):6216. doi: 10.3390/ijms26136216. Int J Mol Sci. 2025. PMID: 40649995 Free PMC article.
References
-
- Slieker R.C., Donnelly L.A., Akalestou E., Lopez-Noriega L., Melhem R., Güneş A., Azar F.A., Efanov A., Georgiadou E., Muniangi-Muhitu H., et al. Identification of biomarkers for glycaemic deterioration in type 2 diabetes. Nat. Commun. 2023;14:2533. doi: 10.1038/s41467-023-38148-7. - DOI - PMC - PubMed
-
- Liu J., Semiz S., van der Lee S.J., van der Spek A., Verhoeven A., van Klinken J.B., Sijbrands E., Harms A.C., Hankemeier T., van Dijk K.W., et al. Metabolomics based markers predict type 2 diabetes in a 14-year follow-up study. Metabolomics. 2017;13:104. doi: 10.1007/s11306-017-1239-2. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

