FLT3-Mutated Leukemic Stem Cells: Mechanisms of Resistance and New Therapeutic Targets
- PMID: 38791898
- PMCID: PMC11119130
- DOI: 10.3390/cancers16101819
FLT3-Mutated Leukemic Stem Cells: Mechanisms of Resistance and New Therapeutic Targets
Abstract
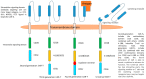
Despite the availability of target drugs in the first and second line, only 30% of FLT3mut AMLs are cured. Among the multiple mechanisms of resistance, those of FLT3mut LSC are the most difficult to eradicate because of their metabolic and genomic characteristics. Reactivation of glycogen synthesis, inhibition of the RAS/MAPK pathway, and degradation of FLT3 may be potential aids to fight the resistance of LSC to FLT3i. LSC is also characterized by the expression of a CD34+/CD25+/CD123+/CD99+ immunophenotype. The receptor and ligand of FLT3, the natural killer group 2 member D ligand (NKGD2L), and CD123 are some of the targets of chimeric antigen receptor T cells (CAR-T), bispecific T-cell engager molecules (BiTEs), CAR-NK and nanoparticles recently designed and reported here. The combination of these new therapeutic options, hopefully in a minimal residual disease (MRD)-driven approach, could provide the future answer to the challenge of treating FLT3mut AML.
Keywords: CAR-NK; CAR-T; FLT3 inhibitors; FLT3-mutated AML; immunotherapy; leukemic stem cell mechanisms of resistance; leukemic stem cell metabolism; leukemic stem cells molecular pathways.
Conflict of interest statement
The author declares no conflict of interest.
Figures
Similar articles
-
Immunoprofiling of leukemic stem cells CD34+/CD38-/CD123+ delineate FLT3/ITD-positive clones.J Hematol Oncol. 2016 Jul 27;9(1):61. doi: 10.1186/s13045-016-0292-z. J Hematol Oncol. 2016. PMID: 27465508 Free PMC article.
-
A novel approach for relapsed/refractory FLT3mut+ acute myeloid leukaemia: synergistic effect of the combination of bispecific FLT3scFv/NKG2D-CAR T cells and gilteritinib.Mol Cancer. 2022 Mar 4;21(1):66. doi: 10.1186/s12943-022-01541-9. Mol Cancer. 2022. PMID: 35246156 Free PMC article.
-
A Leukemia-Associated CD34/CD123/CD25/CD99+ Immunophenotype Identifies FLT3-Mutated Clones in Acute Myeloid Leukemia.Clin Cancer Res. 2015 Sep 1;21(17):3977-85. doi: 10.1158/1078-0432.CCR-14-3186. Epub 2015 May 8. Clin Cancer Res. 2015. PMID: 25957287
-
Overcoming Resistance: FLT3 Inhibitors Past, Present, Future and the Challenge of Cure.Cancers (Basel). 2022 Sep 2;14(17):4315. doi: 10.3390/cancers14174315. Cancers (Basel). 2022. PMID: 36077850 Free PMC article. Review.
-
Paediatric Strategy Forum for medicinal product development of chimeric antigen receptor T-cells in children and adolescents with cancer: ACCELERATE in collaboration with the European Medicines Agency with participation of the Food and Drug Administration.Eur J Cancer. 2022 Jan;160:112-133. doi: 10.1016/j.ejca.2021.10.016. Epub 2021 Nov 25. Eur J Cancer. 2022. PMID: 34840026 Review.
Cited by
-
Tracking Response and Resistance in Acute Myeloid Leukemia through Single-Cell DNA Sequencing Helps Uncover New Therapeutic Targets.Int J Mol Sci. 2024 Sep 17;25(18):10002. doi: 10.3390/ijms251810002. Int J Mol Sci. 2024. PMID: 39337490 Free PMC article.
References
-
- Perl A.E., Larson R.A., Podoltsev N.A., Strickland S., Wang E.S., Atallah E., Schiller G.J., Martinelli G., Neubauer A., Sierra J., et al. Follow-up of patients with R/R FLT3-mutation-positive AML treated with gilteritinib in the phase 3 ADMIRAL trial. Blood. 2022;139:3366–3375. doi: 10.1182/blood.2021011583. - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Miscellaneous