Metagenomic and Antibiotic Resistance Analysis of the Gut Microbiota in Larus relictus and Anatidae Species Inhabiting the Honghaizi Wetland of Ordos, Inner Mongolia, from 2021 to 2023
- PMID: 38792807
- PMCID: PMC11123678
- DOI: 10.3390/microorganisms12050978
Metagenomic and Antibiotic Resistance Analysis of the Gut Microbiota in Larus relictus and Anatidae Species Inhabiting the Honghaizi Wetland of Ordos, Inner Mongolia, from 2021 to 2023
Abstract
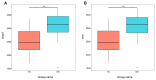
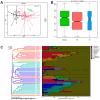
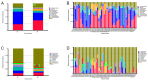
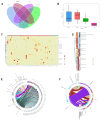
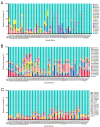
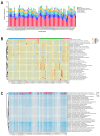
Gut microbes thrive by utilising host energy and, in return, provide valuable benefits, akin to a symbiotic relationship. Here, metagenomic sequencing was performed to characterise and compare the community composition, diversity and antibiotic resistance of the gut microbiota of Relict gull (Larus relictus) and Anatidae species. Alpha diversity analysis revealed that the intestinal microbial richness of L. relictus was significantly lower than that of Anatidae, with distinct differences observed in microbial composition. Notably, the intestines of L. relictus harboured more pathogenic bacteria such as clostridium, which may contribute to the decline in their population and endangered status. A total of 117 strains of Escherichia coli were isolated, with 90.60% exhibiting full susceptibility to 21 antibiotics, while 25.3% exhibited significant biofilm formation. Comprehensive Antibiotic Resistance Database data indicated that glycopeptide resistance genes were the most prevalent type carried by migratory birds, alongside quinolone, tetracycline and lincosamide resistance genes. The abundance of resistance genes carried by migratory birds decreased over time. This metagenomic analysis provides valuable insights into the intestinal microbial composition of these wild bird species, offering important guidance for their conservation efforts, particularly for L. relictus, and contributing to our understanding of pathogen spread and antibiotic-resistant bacteria.
Keywords: Anatidae; Escherichia coli; Larus relictus; biofilm; gut microbiota; metagenomic.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

References
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

